|
[Back to DEMO server]
DEMO Results for job DOT790195786
[Click on DOT790195786_results.tar.bz2 to download the tarball file including all results listed on this page]
|
Query Sequence and Domain Definition
|
>example (262 residues)
|
GIDPNRIVALEWLPVELLLALGIVPYGVADTINYRLWVSEPPLPDSVIDVGLRTEPNLELLTEMKPSFMVWSAGYGPSPEMLARIAPGRGFNFSDGKQPLAMARKSLTEMADLLNLQSAAETHLAQYEDFIRSMKPRFVKRGARPLLLTTLIDPRHMLVFGPNSLFQEILDEYGIPNAWQGETNFWGSTAVSIDRLAAYKDVDVLCFDHDNSKDMDALMATPLWQAMPFVRAGRFQRVPAVWFYGATLSAMHFVRVLDNAIG
|
|
domain 1: 1-94
domain 2: 95-262
|
|
2 domains in total. Different colors represent different domains.
|
|
Input Individual Domain Structures
|
|
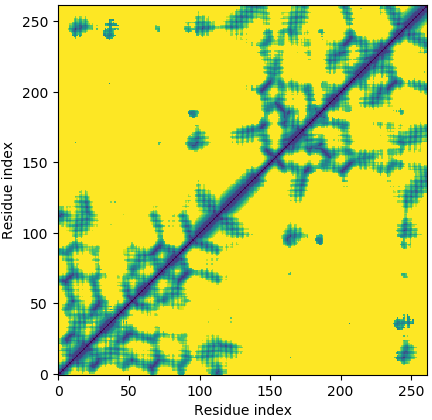
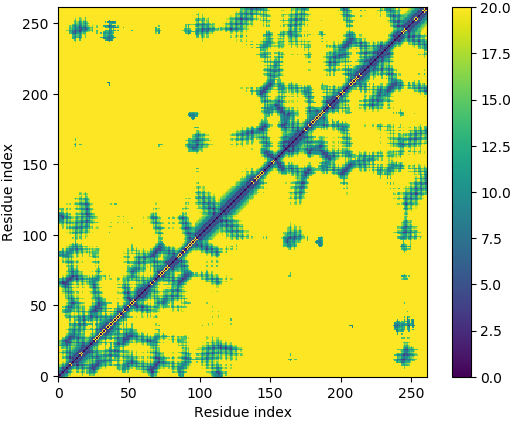
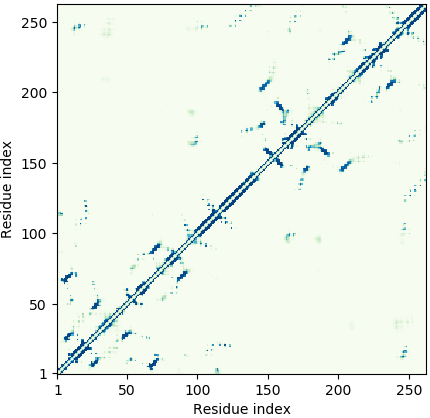
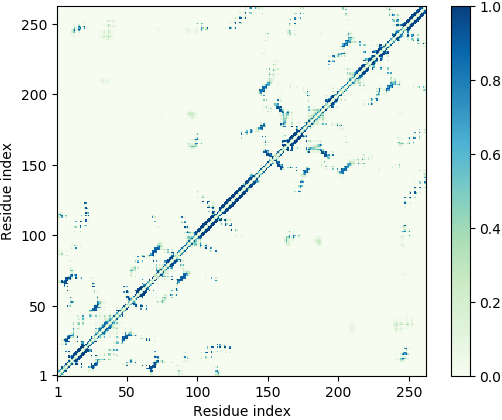
Deep-learning Predicted Distance/Contact Map for Domain Assembly
|
|
Top 10 Full-length Templates for Domain Assembly
|
|
|
Comparison between the best final model and the analogous template
| (a) | Query structure is shown in cartoon, while the structural analogous template is displayed using backbone trace. |
| (c) | TM-scoreh is the score of the template, which is calculated based on harmonic mean of TM-scores between all domain models and the template. |
| (b) | Ranking of proteins is based on the TM-scoreh. |
| (c) | RMSDa is the RMSD between residues that are structurally aligned by TM-align. |
| (d) | IDENb is the percentage sequence identity in the structurally aligned region. |
| (e) | Cov. represents the coverage of the alignment by TM-align and is equal to the number of structurally aligned residues divided by length of the query protein. |
|
|
Final Full-length Models Assembled by DEMO
|
|
|
Top 5 models constructed by DEMO
| (a) | Colored by domain: domain 1 in red; domain 2 in blue. |
| (b) | For each target, DEMO generates an ensemble of structural conformations
by starting from a set of initial models generated by different templates. The server reports up to five final
models sorted by the energy. The accuracy of each model is quantitatively evaluated by estimated TM-score (eTM-score)
and estimated RMSD (eRMSD) that are calculated based on the significance of the structural analogous templates for domain
models assembly, convergence parameters of the domain assembly simulations, satisfaction degrees of the inter-domain
distances/interfaces, and the estimated accuracy of the individual domain model. eTM-score is typically in the range
of [0,1], where an eTM-score of higher value signifies a model with a high confidence and vice-versa. |
| (d) | Since the top 5 models are ranked by the energy,
it is possible that the lower-rank models have a higher eTM-score in rare cases. Although the first model has a better quality
in most cases, it is also possible that the lower-rank models have a better quality than the higher-rank models as seen in
our benchmark tests. |
| More about eTM-score and eRMSD |
|
[Back to DEMO server]
|
Reference:
Xiaogen Zhou, Chunxiang Peng, Wei Zheng, Yang Li, Guijun Zhang, Yang Zhang.
DEMO2: Multi-domain protein structures assembly by coupling quaternary structural
alignment with deep-learning inter-domain restraint prediction.
Nucleic Acids Research, 50(W1): W235-W245 (2022).
Xiaogen Zhou, Jun Hu, Chengxin Zhang, Guijun Zhang, Yang Zhang.
Assembling multidomain protein structures through analogous global structural alignments.
Proceedings of the National Academy of Sciences, 116: 15930-15938 (2019).
|
|