

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MNGVSEGTRGCSDRQPGVLTRDRSCSRKMNSSGCLSEEVGSLRPLTVVILSASIVVGVLGNGLVLWMTVFRMARTVSTVCFFHLALADFMLSLSLPIAMYYIVSRQWLLGEWACKLYITFVFLSYFASNCLLVFISVDRCISVLYPVWALNHRTVQRASWLAFGVWLLAAALCSAHLKFRTTRKWNGCTHCYLAFNSDNETAQIWIEGVVEGHIIGTIGHFLLGFLGPLAIIGTCAHLIRAKLLREGWVHANRPKRLLLVLVSAFFIFWSPFNVVLLVHLWRRVMLKEIYHPRMLLILQASFALGCVNSSLNPFLYVFVGRDFQEKFFQSLTSALARAFGEEEFLSSCPRGNAPRE | |
| CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCSSSSSCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCC | |
| 99988899998989999867888888887877676655314888799999999999999999989988870678688999999999999999989999999805999897289999999999999999999999999989999985301004343468777689999999999899998606844993688324887553103320147789999999999999999999999999999999974787446709999999999999995899999999999874466316899999999999999999979899999438989999999989981687755677788999999999 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | |
| | | | | | | | | | | | | | | | | | | |
| MNGVSEGTRGCSDRQPGVLTRDRSCSRKMNSSGCLSEEVGSLRPLTVVILSASIVVGVLGNGLVLWMTVFRMARTVSTVCFFHLALADFMLSLSLPIAMYYIVSRQWLLGEWACKLYITFVFLSYFASNCLLVFISVDRCISVLYPVWALNHRTVQRASWLAFGVWLLAAALCSAHLKFRTTRKWNGCTHCYLAFNSDNETAQIWIEGVVEGHIIGTIGHFLLGFLGPLAIIGTCAHLIRAKLLREGWVHANRPKRLLLVLVSAFFIFWSPFNVVLLVHLWRRVMLKEIYHPRMLLILQASFALGCVNSSLNPFLYVFVGRDFQEKFFQSLTSALARAFGEEEFLSSCPRGNAPRE | |
| 75442442533433434313333323343344404464241010000221120023133332200000001331100000013311030100000001000002642100100110010113211111000000013102000000020232232210000000011000000100000020254742210102034444334323422220210111111120133002103300010011023244552300000000000001003120100000100230334431330010000001010133033200000000330141014002300441045644444346663468 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCSSSSSCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCC MNGVSEGTRGCSDRQPGVLTRDRSCSRKMNSSGCLSEEVGSLRPLTVVILSASIVVGVLGNGLVLWMTVFRMARTVSTVCFFHLALADFMLSLSLPIAMYYIVSRQWLLGEWACKLYITFVFLSYFASNCLLVFISVDRCISVLYPVWALNHRTVQRASWLAFGVWLLAAALCSAHLKFRTTRKWNGCTHCYLAFNSDNETAQIWIEGVVEGHIIGTIGHFLLGFLGPLAIIGTCAHLIRAKLLREGWVHANRPKRLLLVLVSAFFIFWSPFNVVLLVHLWRRVMLKEIYHPRMLLILQASFALGCVNSSLNPFLYVFVGRDFQEKFFQSLTSALARAFGEEEFLSSCPRGNAPRE | |||||||||||||||||||||||||
| 1 | 4n6hA | 0.29 | 0.28 | 0.90 | 2.96 | Download | LANEGKVKEAQAAAEQLKTTRNAYIQKYLGSPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRYKMKTATNIYIFNLALADALATSTLPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRDGAVVCMLQFPSP----------SWYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLV---DIDRRDPLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKP---------------------- | |||||||||||||||||||
| 2 | 4mbsA | 0.24 | 0.22 | 0.81 | 4.15 | Download | ----------------------------------KINVKQIAARLLPPLYSLVFIFGFVGNMLVILILIYKRLKSMTDIYLLNLAISDLFFLLTVPFWAHYAA-AQWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEGLHYTCSSHFPYSQ---------YQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLREEEKKRHRDVRLIFTIMIVYFLFWAPYNIVLLLNTFQEFLNNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------ | |||||||||||||||||||
| 3 | 4n6hA | 0.31 | 0.27 | 0.82 | 3.91 | Download | ------------------------------SPGARSASSLALAIAITALYSAVCAVGLLGNVLVMFGIVRTKMKTATNIYIFNLALADALATSTLPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRDGAVVCMLQFPSPS----------WYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSVRLLSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIDRRD---PLVVAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG-------------------- | |||||||||||||||||||
| 4 | 4djh | 0.29 | 0.26 | 0.77 | 1.52 | Download | --------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRTKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVDVDVIECSLQFPDDDYS-----------WWLFMICVFIFAFVIPVLIIIVCYTLMILRLKSVRLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA---------ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----------------------- | |||||||||||||||||||
| 5 | 4djh | 0.29 | 0.26 | 0.78 | 1.17 | Download | --------------------------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRTKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVRVDVIECSLQFPDDDY---------SWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGS---A------ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP----------------------- | |||||||||||||||||||
| 6 | 5o9hA | 0.28 | 0.27 | 0.81 | 3.30 | Download | -------------------------------------TLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVQHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEYPKVLCGVDHD-------------KRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFIFWLPYQVTGIMMSFLE--PSSPTFLLLKKLDSLCVSFAYINCCINPIIYVVAGQGFQKSLPELLREVLTEESVVR-------------- | |||||||||||||||||||
| 7 | 4djh | 0.29 | 0.26 | 0.77 | 1.74 | Download | ---------------------------------------PAIPVIITAVYSVVFVVGLVGNSLVMFVIIRTKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVDVDVIECSLQFPDDDYSWWDL---------FMKICVFIFAFVIPVLIIIVCYTLMILRLKREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSA---------ALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF------------------------ | |||||||||||||||||||
| 8 | 4mbsA | 0.24 | 0.22 | 0.81 | 4.64 | Download | -------------------------------PCQKINVKQIAARLLPPLYSLVFIFGFVGNMLVILILINYKRKSMTDIYLLNLAISDLFFLLTVPFWAHYAAA-QWDFGNTMCQLLTGLYFIGFFSGIFFIILLTIDRYLAVVHAVFALKARTVTFGVVTSVITWVVAVFASLPNIIFTRSQKEGLHYTCSSHFPYSQ---------YQFWKNFQTLKIVILGLVLPLLVMVICYSGILKTLLREKKRHRD--VRLIFTIMIVYFLFWAPYNIVLLLNTFQEFFGNCSSSNRLDQAMQVTETLGMTHCCINPIIYAFVGEEFRNYLLVFFQ------------------------ | |||||||||||||||||||
| 9 | 5o9hA | 0.27 | 0.27 | 0.82 | 3.60 | Download | ------------------------------------NTLRVPDILALVIFAVVFLVGVLGNALVVWVTAFEAKRTINAIWFLNLAVADFLACLALPALFTSIVHHHWPFGGAACSILPSLILLNMYASILLLATISADRFLLVFKPAWCQRFRGAGLAWILCAVAWGLALLLTIPSALYRVVREEPPKVLCGVDHD-------------KRRERAVAIVRLVLGFLWPLLTLTICYTFILLRTWSARETRSTKTLKVVVAVVASFFIFWLPYQVTGIMMSFL--EPSSPTFLLLKKLDSLCVSFAYINCCINPIIYVVAGQGFQKSLPELLREVLTEESVVR-------------- | |||||||||||||||||||
| 10 | 5c1mA | 0.28 | 0.24 | 0.80 | 4.56 | Download | ---------------------------GSHSLPQTGSPSMVTAITIMALYSIVCVVGLFGNFLVMYVIVYTKMKTATNIYIFNLALADALATSTLPFQSVNYLMGTWPFGNILCKIVISIDYYNMFTSIFTLCTMSVDRYIAVCHPVKALDFRTPRNAKIVNVCNWILSSAIGLPVMFMATTKYRQGSIDCTLTFSHP----------TWYWENLLKICVFIFAFIMPVLIITVCYGLMILRLKSVRMRNLRRITRMVLVVVAVFIVCWTPIHIYVIIKALITIPET----TFQTVSWHFCIALGYTNSCLNPVLYAFLDENFKRCF----------------------------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
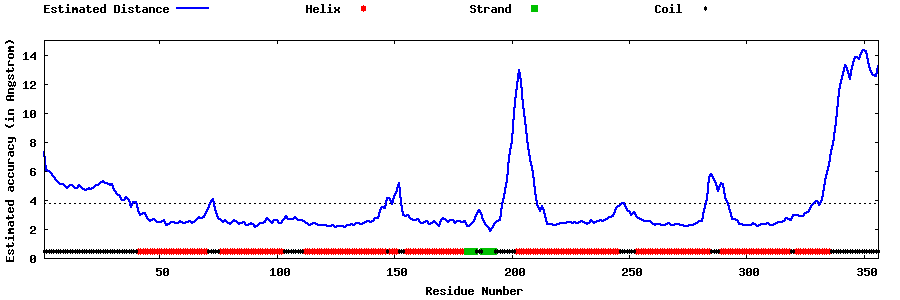
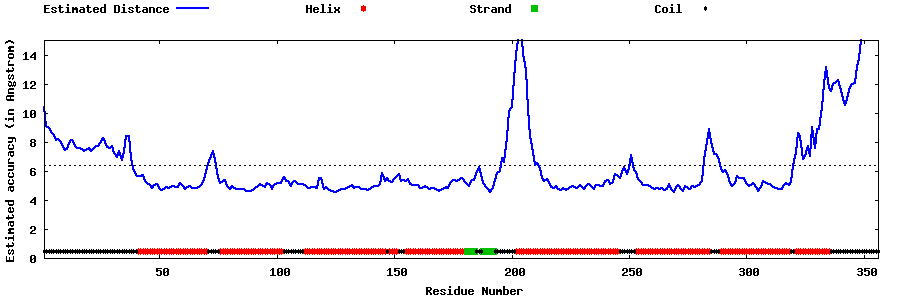
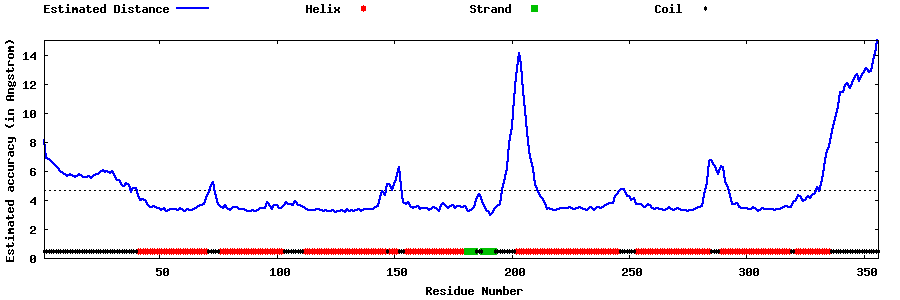
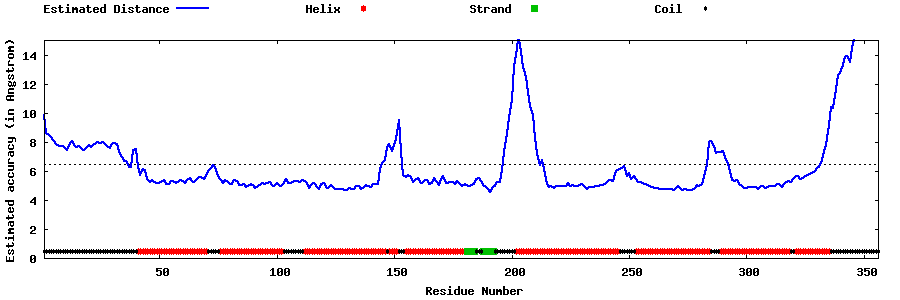
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||