

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MDFLNSSDQNLTSEELLNRMPSKILVSLTLSGLALMTTTINSLVIAAIIVTRKLHHPANYLICSLAVTDFLVAVLVMPFSIVYIVRESWIMGQVVCDIWLSVDITCCTCSILHLSAIALDRYRAITDAVEYARKRTPKHAGIMITIVWIISVFISMPPLFWRHQGTSRDDECIIKHDHIVSTIYSTFGAFYIPLALILILYYKIYRAAKTLYHKRQASRIAKEEVNGQVLLESGEKSTKSVSTSYVLEKSLSDPSTDFDKIHSTVRSLRSEFKHEKSWRRQKISGTRERKAATTLGLILGAFVICWLPFFVKELVVNVCDKCKISEEMSNFLAWLGYLNSLINPLIYTIFNEDFKKAFQKLVRCRC | |
| CCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCC | |
| 998768999988787777728999999999999999999999850001302577764479999999999999999874999999828056886899999999999999999999999986578535342187426489999999999999999999999848776888982798756669999999999999999999999999999999987752134453323333322122333334444333334443332334432122333322234322110356777778877888888998898882008899999987167767369999999999999717689898187999999999807899 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | |
| | | | | | | | | | | | | | | | | | | | |
| MDFLNSSDQNLTSEELLNRMPSKILVSLTLSGLALMTTTINSLVIAAIIVTRKLHHPANYLICSLAVTDFLVAVLVMPFSIVYIVRESWIMGQVVCDIWLSVDITCCTCSILHLSAIALDRYRAITDAVEYARKRTPKHAGIMITIVWIISVFISMPPLFWRHQGTSRDDECIIKHDHIVSTIYSTFGAFYIPLALILILYYKIYRAAKTLYHKRQASRIAKEEVNGQVLLESGEKSTKSVSTSYVLEKSLSDPSTDFDKIHSTVRSLRSEFKHEKSWRRQKISGTRERKAATTLGLILGAFVICWLPFFVKELVVNVCDKCKISEEMSNFLAWLGYLNSLINPLIYTIFNEDFKKAFQKLVRCRC | |
| 344313233324444333310110000021132033033112000000012330220000000000100110010002300000012302103100010000001000000200000010001000200404342232000000000022022002000001233354542020323310000000210131001000200120020024214423444444433444333433343344334323342332333343432333233233323443333332232244333220000001111000221011000100034130241000000120120011000000100430140023003032 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 320 340 360 | | | | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHCCHHHHHHHHHHHCCCC MDFLNSSDQNLTSEELLNRMPSKILVSLTLSGLALMTTTINSLVIAAIIVTRKLHHPANYLICSLAVTDFLVAVLVMPFSIVYIVRESWIMGQVVCDIWLSVDITCCTCSILHLSAIALDRYRAITDAVEYARKRTPKHAGIMITIVWIISVFISMPPLFWRHQGTSRDDECIIKHDHIVSTIYSTFGAFYIPLALILILYYKIYRAAKTLYHKRQASRIAKEEVNGQVLLESGEKSTKSVSTSYVLEKSLSDPSTDFDKIHSTVRSLRSEFKHEKSWRRQKISGTRERKAATTLGLILGAFVICWLPFFVKELVVNVCDKCKISEEMSNFLAWLGYLNSLINPLIYTIFNEDFKKAFQKLVRCRC | |||||||||||||||||||||||||
| 1 | 4iaqA | 0.46 | 0.47 | 0.94 | 2.70 | Download | -----------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWRQASE-----CVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLEDNWETLNDNLKVINAAQVKDALTKMRAAALDAQKATPDSPEMKDFRHGFDILVGQIDDNAYIQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLVMP------IHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK- | |||||||||||||||||||
| 2 | 5uenA | 0.21 | 0.24 | 0.94 | 4.79 | Download | --------------------AFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQTYFH--TCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSAVERAWAAAISMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNWETLNDNVKDALTKMRAAALDAPEMKDFALKLANEGKVKEAQAAAEQLKTTRNAYIQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHF | |||||||||||||||||||
| 3 | 5uenA | 0.22 | 0.24 | 0.93 | 3.57 | Download | -----------------SISAFQAAYIGIEVLIALVSVPGNVLVIWAVKVNQALRDATFCFIVSLAVADVAVGALVIPLAILINIGPQTYF--HTCLMVACPVLILTQSSILALLAIAVDRYLRVKIPLRYKMVVTPRRAAVAIAGCWILSFVVGLTPMFGWNNLSGSMGEPVIKCEMEYMVYFNFFVWVLPPLLLMVLIYLEVFYLIRKQLADLEDNWETLNDNVKMRAAALDAPEMKDFALKLANEGKVKEAQAAAEQLKTTRNAY-----IQKYLERARSTLQKELKIAKSLALILFLFALSWLPLHILNCITLFCPSCHKPSILTYIAIFLTHGNSAMNPIVYAFRIQKFRVTFLKIWNDHF | |||||||||||||||||||
| 4 | 4ib4 | 0.25 | 0.29 | 0.96 | 1.57 | Download | -------------EEQGNK---LHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIETNPNNITCVLTKERGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQRAAALDAQKFRHGFDGQIDALKLANEGKVKEAQKTTRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNY | |||||||||||||||||||
| 5 | 3uon | 0.29 | 0.33 | 0.94 | 1.24 | Download | -------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFITVEDGECYIQFSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINIFEMLRIDEGLRLKESPSLNAAKLDKAIGRNTNGVITKNAKLQQKRWDENLAKSRNQTPTFRTGTWDAYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPC-IPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM--- | |||||||||||||||||||
| 6 | 4iaqA | 0.45 | 0.47 | 0.93 | 3.39 | Download | ------------IYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWRQA-----SECVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLEDNWETLNDNLKVIDSPEMKDFRHGFDILVGQIDDALKLANEGKVKEAQAAAEQLKTTRNAYIQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLVMPI------HLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK- | |||||||||||||||||||
| 7 | 4ib4 | 0.25 | 0.29 | 0.93 | 1.77 | Download | ----------------------LHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMEAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVIKGIETN-PNNITCVLTKERGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWETLNDNLKVIEKADNAAQVTRAAALDAQKKGFDILVGANEGKVKEAQAAAEQLKRNAYIQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNTLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCN- | |||||||||||||||||||
| 8 | 2rh1A | 0.30 | 0.33 | 0.92 | 4.64 | Download | ------------------DEVWVVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRATHEETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKRQLNIFEMLRIDEGVAGFTNSLRMLQQKRWDEAAVNLAKSRWYNQTPNRAKRVITTFRTGTWDAYKFCL--------KEHKALKTLGIIMGTFTLCWLPFFIVNIVHVIQDNL-IRKEVYILLNWIGYVNSGFNPLIYC-RSPDFRIAFQELLC--L | |||||||||||||||||||
| 9 | 4iaqA | 0.45 | 0.47 | 0.94 | 3.03 | Download | -----------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWRQASE-----CVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLEDNWETLNDNLKVINAAQVKDALTKMRAAALDAQKATPDSPEGKVKEAQAAAEQLKTTRNAYIQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLVMP------IHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK- | |||||||||||||||||||
| 10 | 4iaqA | 0.46 | 0.47 | 0.94 | 3.19 | Download | -----------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFWR-----QASECVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLEDNWETLNDNLKVINAAQVKDALTKMRAAALDAQKATPDSPEGKVKEAQAAAEQLKTTRNAYIQKYLLMAARERKATKTLGIILGAFIVCWLPFFIISLV------MPIHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK- | |||||||||||||||||||
| ||||||||||||||||||||||||||
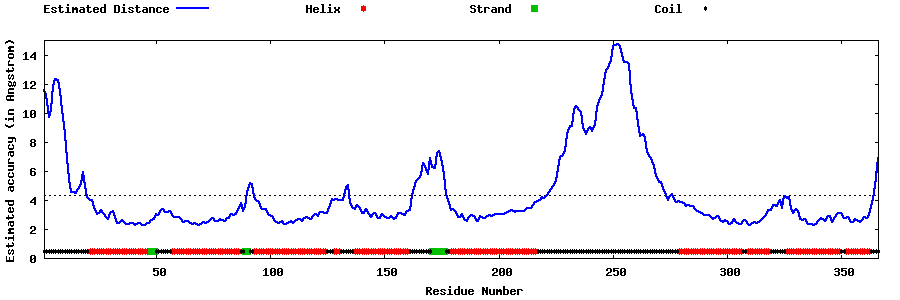
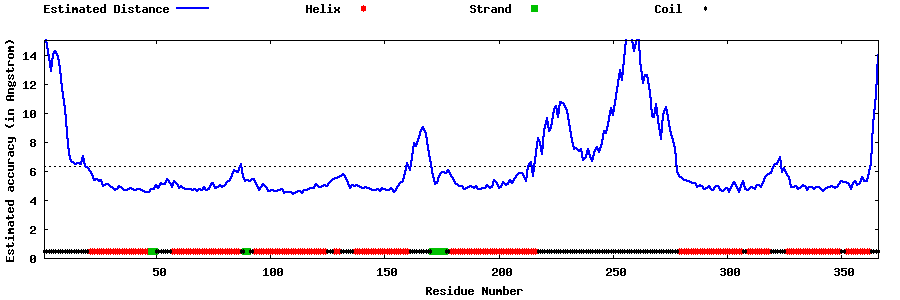
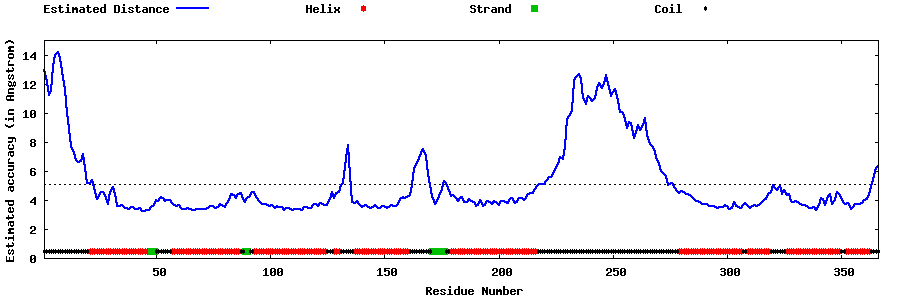
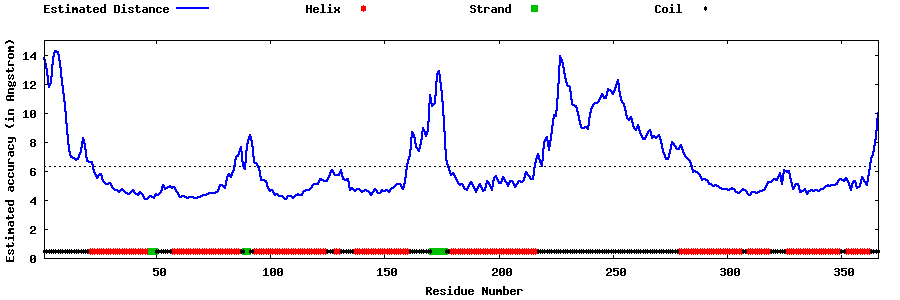
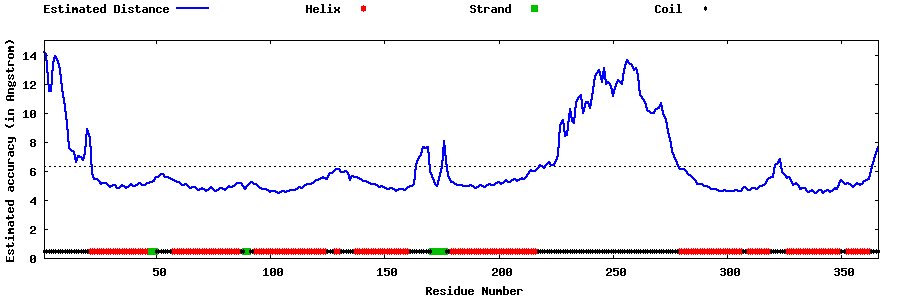
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||