

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MQAALTAFFVLLFSLLSLLGIAANGFIVLVLGREWLRYGRLLPLDMILISLGASRFCLQLVGTVHNFYYSAQKVEYSGGLGRQFFHLHWHFLNSATFWFCSWLSVLFCVKIANITHSTFLWLKWRFPGWVPWLLLGSVLISFIITLLFFWVNYPVYQEFLIRKFSGNMTYKWNTRIETYYFPSLKLVIWSIPFSVFLVSIMLLINSLRRHTQRMQHNGHSLQDPSTQAHTRALKSLISFLILYALSFLSLIIDAAKFISMQNDFYWPWQIAVYLCISVHPFILIFSNLKLRSVFSQLLLLARGFWVA | |
| CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCHHHHHHHHHHHCCHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCCCCC | |
| 9748999999999999999999999999999999980798886779999999999999999987066032153211551799999999999939999999999999835641789779999998622309999999999999999999974012013420246787068874230689999999999999999999999999999999999986188999999980899999999999999999999999999999844453999999999999788858999618749999999999865570039 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MQAALTAFFVLLFSLLSLLGIAANGFIVLVLGREWLRYGRLLPLDMILISLGASRFCLQLVGTVHNFYYSAQKVEYSGGLGRQFFHLHWHFLNSATFWFCSWLSVLFCVKIANITHSTFLWLKWRFPGWVPWLLLGSVLISFIITLLFFWVNYPVYQEFLIRKFSGNMTYKWNTRIETYYFPSLKLVIWSIPFSVFLVSIMLLINSLRRHTQRMQHNGHSLQDPSTQAHTRALKSLISFLILYALSFLSLIIDAAKFISMQNDFYWPWQIAVYLCISVHPFILIFSNLKLRSVFSQLLLLARGFWVA | |
| 7433233311332331233133202200210111004434031011000000211010010133310000000321333210000101123312300100000010000000213110000013304310011023123333311211113233233333344341100131314332201111333333333331332332011002202430343353441231410220010012022223323311321111223342201021113002102300000021143014001300210210018 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSSCCHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCHHHHHHHHHHHCCHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCSSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCCCCC MQAALTAFFVLLFSLLSLLGIAANGFIVLVLGREWLRYGRLLPLDMILISLGASRFCLQLVGTVHNFYYSAQKVEYSGGLGRQFFHLHWHFLNSATFWFCSWLSVLFCVKIANITHSTFLWLKWRFPGWVPWLLLGSVLISFIITLLFFWVNYPVYQEFLIRKFSGNMTYKWNTRIETYYFPSLKLVIWSIPFSVFLVSIMLLINSLRRHTQRMQHNGHSLQDPSTQAHTRALKSLISFLILYALSFLSLIIDAAKFISMQNDFYWPWQIAVYLCISVHPFILIFSNLKLRSVFSQLLLLARGFWVA | |||||||||||||||||||||||||
| 1 | 4djhA | 0.11 | 0.18 | 0.90 | 1.36 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMN-SWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVRE---DVDVIECSLQFYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSDR-----------NLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------- | |||||||||||||||||||
| 2 | 5tjvA | 0.11 | 0.20 | 0.89 | 2.75 | Download | LNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHS--RSLRCRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRKDSRN-VFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLA----YKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVCSDIFPHID---------ETYLMFWIGVTSVLLLFIVYAYMYILWKAGKRAMSFSD--------QARMDIRLAKTLVLILVVLIICWGPLLAIMVYDVFGKMTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 3 | 4n6hA | 0.11 | 0.20 | 0.92 | 2.14 | Download | SSLALAIAITALYSAVCAVGLLGNVLVMFGIVRY--TKMK-TATNIYIFNLALADALATST--LPFQSAKYLMTWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPAKAKLINICIWVLASGVGVPIMVMA-------VTRPRDGAVVCMLQPSWYWDTVTKICVFLFVVPILIITVCYGLMLLRLRSV-----RLLSGSK-EKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---- | |||||||||||||||||||
| 4 | 4djh | 0.10 | 0.16 | 0.95 | 1.56 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR---YTKMKTATNIYIFNLALADALVTTT-MPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKAL----DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDIECSLQFPDDDYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNIFTPAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEAGSAALSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------- | |||||||||||||||||||
| 5 | 5glh | 0.11 | 0.20 | 0.90 | 1.19 | Download | IKETFKYINTVVSCLVFVLGIIGNSTLLYIIYK--------NGPNILIASLALGDLLHIVIAIPINVYKLLAEDWPFGAEMCKLVPFIQKASVGITVLSLCALSIDRYRAVASW--------------WTAVEIVLIWVVSVVLAVPEAIGFDIITMD-YKGSYLRICLLHPVQKAFMQFYWWLFSFYFCLPLAITAFFYTLMTCEMLRKNEGLRLTWDAYLNDHLKQRREVAKTVFCLVLVFALCWLPLHLARILKTLYNNVLDYIGINMASLNSCANPIALYLVSKRFKNAFKSAL--------- | |||||||||||||||||||
| 6 | 4djhA | 0.11 | 0.18 | 0.90 | 1.51 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRY---TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMN-SWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVRE---DVDVIECSLQFPSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSD-----------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------- | |||||||||||||||||||
| 7 | 3uon | 0.11 | 0.18 | 0.92 | 1.72 | Download | -----VVFIVLVAGSLSLVTIIGNILVMVSIKV---NRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIVGVRT--VEDGECYIQ----FFS--NAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINIFEADAYPPPSREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM-------- | |||||||||||||||||||
| 8 | 4n6hA | 0.12 | 0.20 | 0.92 | 2.92 | Download | SSLALAIAITALYSAVCAVGLLGNVLVMFGIVRY---TKMKTATNIYIFNLALADALATST-LPFQSAKYLMETWPFGELLCKAVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVK----ALDFRTPAKAKLINICIWVLASGVGVPIMVMAVTRPRD-------GAVVCMLQFYWDTVTKICVFLFAFVVPILIITVCYGLMLLRLRSV------RLLSGSKEKDRSLRRITRMVLVVVGAFVVCWAPIHIFVIVWTLVDIAALHLCIALGYANSSLNPVLYAFLDENFKRCFRQLCRKPCG---- | |||||||||||||||||||
| 9 | 4djhA | 0.11 | 0.18 | 0.90 | 1.47 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMN-SWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIE---CSLQFPDDWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLL------------DRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSASSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------- | |||||||||||||||||||
| 10 | 1l9hA | 0.14 | 0.18 | 0.93 | 1.71 | Download | AEPWQFSMLAAYMFLLIMLGFPINFLTLYVTVQ----HKKLTPLNYILLNLAVADLFMVFGGFTTTLYTSLHGYFVFGPTGCNLEGFFATLGGEIALWSLVVLAIERYVVVCPMSNFRFG------ENHAIMGVAFTWVMALACAAPPLVGWSRYIPEGMQCSCGIDYYTPHEETNNESFVIYMFVVHFIIPLIVIFFCYGQLVF----------TVKEAAAATTQKAEKEVTRMVIIMVIAFLICWLPYAGVAFYIFTHQGIFMTIPAFFAKTSAVYNPVIYIMMNKQFRNCMVTTLCCGKNPLGD | |||||||||||||||||||
| ||||||||||||||||||||||||||
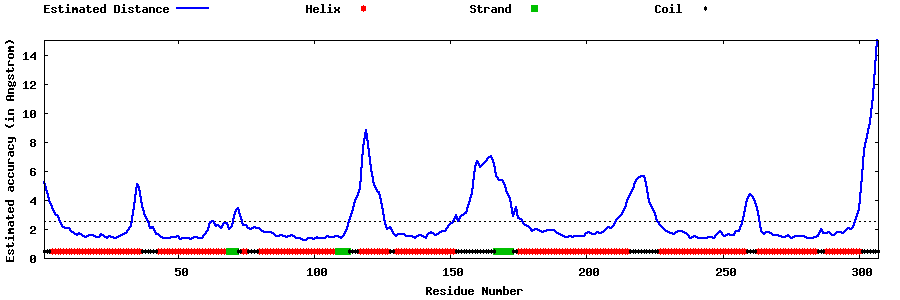
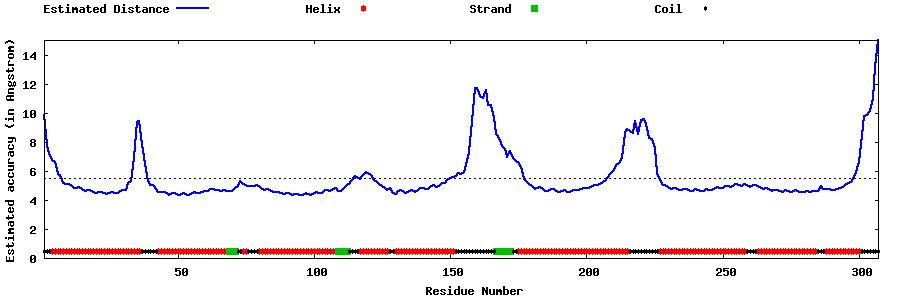
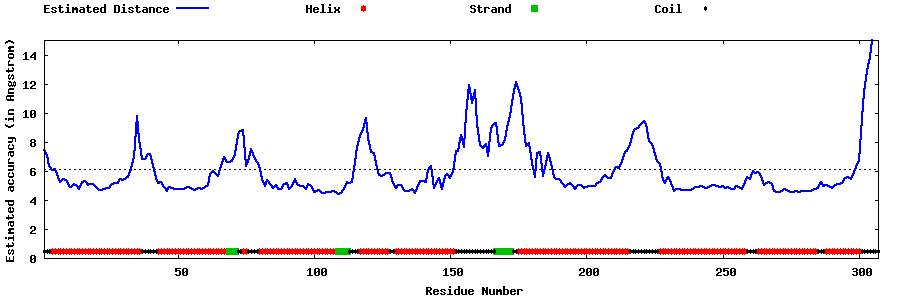
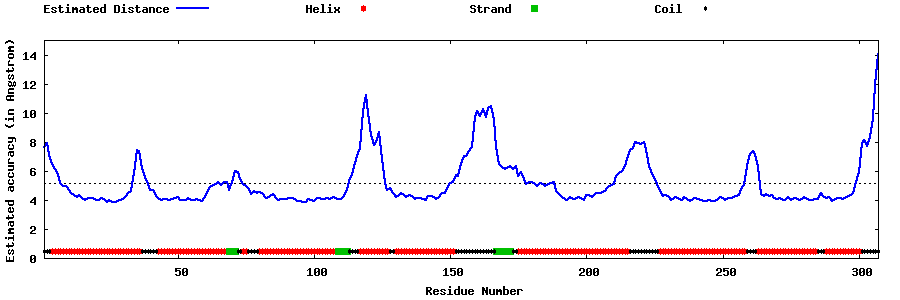
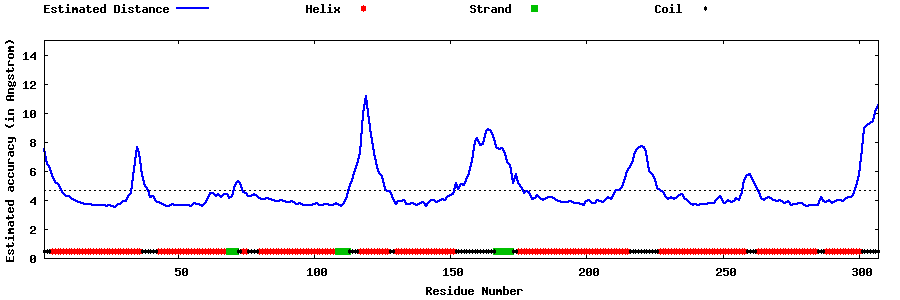
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||