

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTTFIPIIFSSVVVVLFVIGNFANGFIALVNSIERVKRQKISFADQILTALAVSRVGLLWVLLLNWYSTVFNPAFYSVEVRTTAYNVWAVTGHFSNWLATSLSIFYLLKIANFSNLIFLHLKRRVKSVILVMLLGPLLFLACQLFVINMKEIVRTKEYEGNLTWKIKLRSAVYLSDATVTTLGNLVPFTLTLLCFLLLICSLCKHLKKMQLHGKGSQDPSTKVHIKALQTVIFFLLLCAVYFLSIMISVWSFGSLENKPVFMFCKAIRFSYPSIHPFILIWGNKKLKQTFLSVLRQVRYWVKGEKPSSP | |
| CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCSSCCCCCCCC | |
| 986899999999999999999999999999999998279887787999999999999999999999997635452156779678779999838999999999999844660899779999998466608999999999999999999713330047888850377652226899999999999999999999999999999999999860889999999818999999999999999999999999999998505771999999999998378866988418669999999999863121368799997 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTTFIPIIFSSVVVVLFVIGNFANGFIALVNSIERVKRQKISFADQILTALAVSRVGLLWVLLLNWYSTVFNPAFYSVEVRTTAYNVWAVTGHFSNWLATSLSIFYLLKIANFSNLIFLHLKRRVKSVILVMLLGPLLFLACQLFVINMKEIVRTKEYEGNLTWKIKLRSAVYLSDATVTTLGNLVPFTLTLLCFLLLICSLCKHLKKMQLHGKGSQDPSTKVHIKALQTVIFFLLLCAVYFLSIMISVWSFGSLENKPVFMFCKAIRFSYPSIHPFILIWGNKKLKQTFLSVLRQVRYWVKGEKPSSP | |
| 733333331133133123313321220021001100443403201100000021100000113330000001120233300000001012212300100000010000000223110000012304310011012122323321131303221325434212123131332210111123333333233113323330220022023203433444411314002200100130221023113200200011233433211110112102101200101003143013001300220101036463668 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCHHHCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSSSCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCSSSSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHCCCSSCCCCCCCC MTTFIPIIFSSVVVVLFVIGNFANGFIALVNSIERVKRQKISFADQILTALAVSRVGLLWVLLLNWYSTVFNPAFYSVEVRTTAYNVWAVTGHFSNWLATSLSIFYLLKIANFSNLIFLHLKRRVKSVILVMLLGPLLFLACQLFVINMKEIVRTKEYEGNLTWKIKLRSAVYLSDATVTTLGNLVPFTLTLLCFLLLICSLCKHLKKMQLHGKGSQDPSTKVHIKALQTVIFFLLLCAVYFLSIMISVWSFGSLENKPVFMFCKAIRFSYPSIHPFILIWGNKKLKQTFLSVLRQVRYWVKGEKPSSP | |||||||||||||||||||||||||
| 1 | 4djhA | 0.14 | 0.19 | 0.89 | 1.46 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKA----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPDWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSDR-----------NLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF-------------P | |||||||||||||||||||
| 2 | 5tjvA | 0.10 | 0.20 | 0.89 | 2.76 | Download | LNPSQQLAIAVLSLTLGTFTVLENLLVLCVILH--SRSLRCRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLA----YKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVCSDIF-----PHIDETYLMFWIGVTSVLLLFIVYAYMYILWKAGKRAMSFSD--------QARMDIRLAKTLVLILVVLIICWGPLLAIMVYDVFGKMNTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------------- | |||||||||||||||||||
| 3 | 4djhA | 0.13 | 0.19 | 0.89 | 2.19 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRY--TKMK-TATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKAL----DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLL-------------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------- | |||||||||||||||||||
| 4 | 4ib4 | 0.11 | 0.21 | 0.92 | 1.57 | Download | EQGNKLHWAALLILMVIIPTIGGNTLVILAVSLE---KKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFEMWPLPLVLCAWLFLDVLFSTASIWHLCAISVDRYIAIK---KPIQAN-QYNSRATAFIKITVVWLIIGIAIPVPIGIETNPNNTCVLTKE-----RFGDFML--FGSLAAFFTPLAIMIVTYFLTIHALQKKAADLEDNWEQKYLQTISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLDNLQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR---------- | |||||||||||||||||||
| 5 | 3odu | 0.16 | 0.25 | 0.96 | 1.27 | Download | NANFNKIFLPTIYSIIFLTGIVGNGLVILVMG----YQKKRSMTDKYRLHLSVADLLFVIT-LPFWAVDAVANWYFGNFLCKAVHVIYTVNLYSSVWILAFISLDRYLAIVHATNSQ----RPRKLLAEKVVYVGVWIPALLLTIPDFIFANVSEADDRYICDRFYPNDLWVVVFQFQHIMVGLILPGIVILSCYCIIISKLSHSGSNIFEMLRDAYGSKGHQKRKALKTTVILILAFFACWLPYYIGISIDFILLVHKWISITEALAFFHCCLNPILYAFLGAKFKTSAQHALTSGRPLEVLFQ---- | |||||||||||||||||||
| 6 | 4djhA | 0.13 | 0.19 | 0.90 | 1.65 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRY---TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKAL----DFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSD-----------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------------- | |||||||||||||||||||
| 7 | 4djh | 0.13 | 0.16 | 0.92 | 1.71 | Download | --PAIPVIITAVYSVVFVVGLVGNSLVMFVIIR---YTKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIE--CSLQFPDDDWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSGNAYREKDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGS---AALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF-------------- | |||||||||||||||||||
| 8 | 4buoA | 0.16 | 0.20 | 0.91 | 2.38 | Download | TDIYSKVLVTAIYLALFVVGTVGNSVTLFTLAR---KKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWPWAFGDAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPGGLVCTPIVTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQ---------PGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDEYFYMLTNALVYVSAAINPILYNLVSANFRQVFLSTL--------------- | |||||||||||||||||||
| 9 | 4djhA | 0.14 | 0.19 | 0.89 | 1.48 | Download | -SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIR----TKMKTATNIYIFNLALADALVTTTMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCPVKA-----LDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVREDVDVIECSLQFPWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLSD-----------RNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAASSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCF-------------P | |||||||||||||||||||
| 10 | 1l9hA | 0.12 | 0.19 | 0.94 | 1.53 | Download | AEPWQFSMLAAYMFLLIMLGFPINFLTLYVTVQ----HKKLTPLNYILLNLAVADLFMVFGGFTTTLYTSLHGYFFGPTGCNLEGFFATLGGEIALWSLVVLAIERYVVVCPMSNFRF------GENHAIMGVAFTWVMALACAAPPLVGWSRYIPEGMQCSCGIDYYTPHNESFVIYMFVVHFIIPLIVIFFCYGQLVF----------TVKEAAAATTQKAEKEVTRMVIIMVIAFLICWLPYAGVAFYIFTHQGPIFMTIPAFFAKTSAVYNPVIYIMMNKQFRNCMVTTLCCGKNPLGDSTTVSK | |||||||||||||||||||
| ||||||||||||||||||||||||||
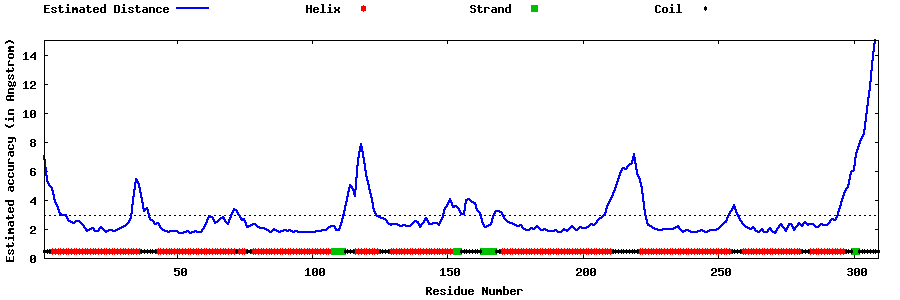
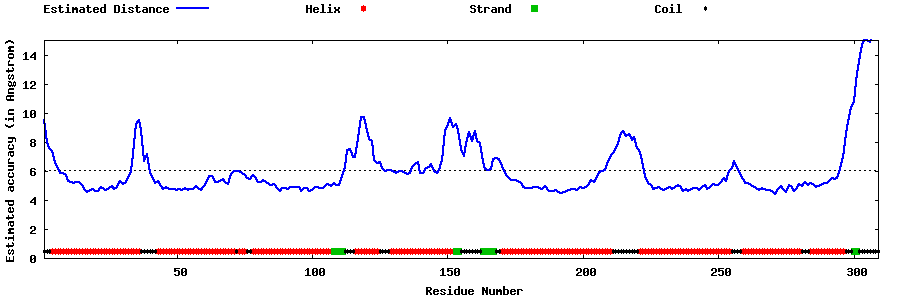
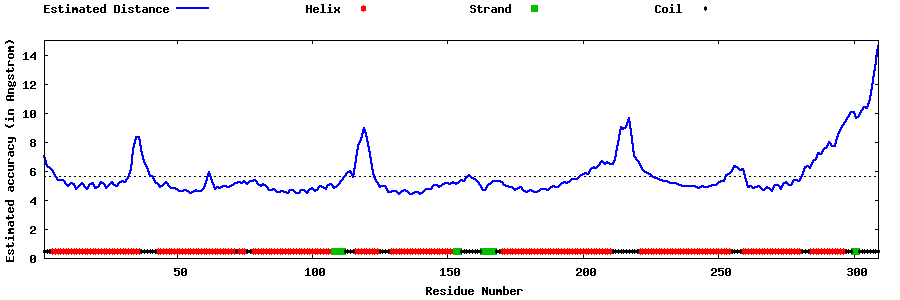
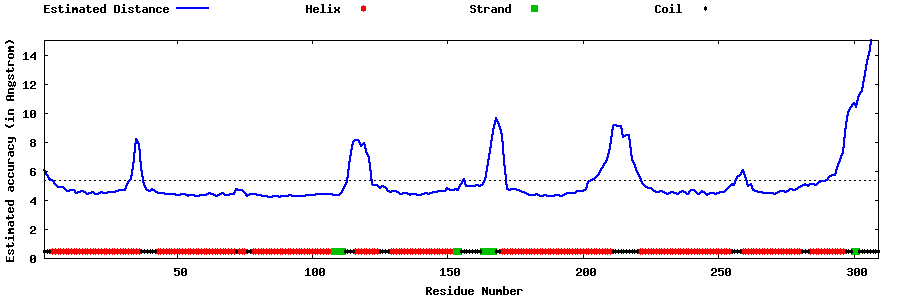
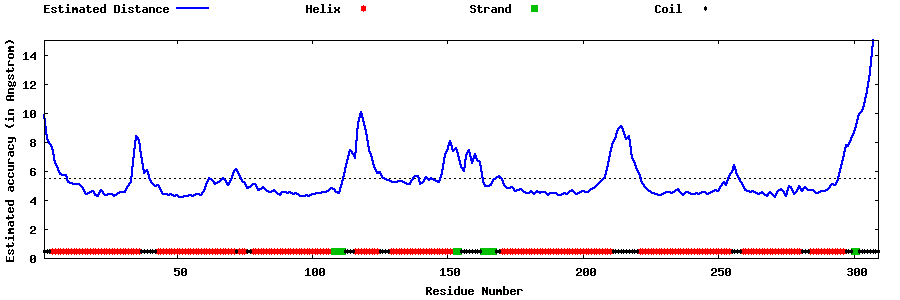
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||