

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MGPRNQTAVSEFLLMKVTEDPELKLIPFSLFLSMYLVTILGNLLILLAVISDSHLHTPMYFLLFNLSFTDICLTTTTVPKILVNIQAQNQSITYTGCLTQICLVLVFAGLESCFLAVMAYDRYVAICHPLRYTVLMNVHFWGLLILLSMFMSTMDALVQSLMVLQLSFCKNVEIPLFFCEVVQVIKLACSDTLINNILIYFASSVFGAIPLSGIIFSYSQIVTSVLRMPSARGKYKAFSTCGCHLSVFSLFYGTAFGVYISSAVAESSRITAVASVMYTVVPQMMNPFIYSLRNKEMKKALRKLIGRLFPF | |
| CCCCCCCCCHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSCCCCCCCCCCCCSSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHHCCCCC | |
| 99988850765630079988426789999999999999998999999997188755639999888999865334148899989983489657378889999999999999999999998750550176543774217608999999999999999999999997517899995589357879987560644389999999999999998699999999999988401457788676204626688989998580541560689878898883899830063625445642466088999999998263899 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MGPRNQTAVSEFLLMKVTEDPELKLIPFSLFLSMYLVTILGNLLILLAVISDSHLHTPMYFLLFNLSFTDICLTTTTVPKILVNIQAQNQSITYTGCLTQICLVLVFAGLESCFLAVMAYDRYVAICHPLRYTVLMNVHFWGLLILLSMFMSTMDALVQSLMVLQLSFCKNVEIPLFFCEVVQVIKLACSDTLINNILIYFASSVFGAIPLSGIIFSYSQIVTSVLRMPSARGKYKAFSTCGCHLSVFSLFYGTAFGVYISSAVAESSRITAVASVMYTVVPQMMNPFIYSLRNKEMKKALRKLIGRLFPF | |
| 85672312002000010154252110101211301220133221000001001300000000000101101102002203001001142330303000001100010022010000000110000000002000000020000001201210010011000000000103624010000123200300011121010101221333033133003201010000001021462321000000000000001110020000001213144322000000013223300100002032023003301433356 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCHHHHHCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHCCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCSSCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCSSCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCSSSCCHHHHHHHHHHHCCHHSSSCCCCCCCCCCCCSSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHHCCCCC MGPRNQTAVSEFLLMKVTEDPELKLIPFSLFLSMYLVTILGNLLILLAVISDSHLHTPMYFLLFNLSFTDICLTTTTVPKILVNIQAQNQSITYTGCLTQICLVLVFAGLESCFLAVMAYDRYVAICHPLRYTVLMNVHFWGLLILLSMFMSTMDALVQSLMVLQLSFCKNVEIPLFFCEVVQVIKLACSDTLINNILIYFASSVFGAIPLSGIIFSYSQIVTSVLRMPSARGKYKAFSTCGCHLSVFSLFYGTAFGVYISSAVAESSRITAVASVMYTVVPQMMNPFIYSLRNKEMKKALRKLIGRLFPF | |||||||||||||||||||||||||
| 1 | 3emlA | 0.22 | 0.24 | 0.89 | 3.59 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAII-VGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLR | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.23 | 0.90 | 2.13 | Download | -RGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFID-FHVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-----------------GWNCEKLQSVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAMLCLLNSTVNPIIYALRSKDLRHAFRSM------- | |||||||||||||||||||
| 3 | 4iaqA | 0.18 | 0.24 | 0.86 | 2.19 | Download | ------------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISL---------PPCVVNTDH--------------------ILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIIQKYLLRERKATKTLGIILGAFIVCWLPFFIISLVMPIHLAIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK--- | |||||||||||||||||||
| 4 | 4djh | 0.15 | 0.26 | 0.90 | 1.56 | Download | -------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP---- | |||||||||||||||||||
| 5 | 4yay | 0.16 | 0.23 | 0.91 | 1.25 | Download | KTTRN-AYIQKYLILNSSDCPYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNV-FF------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIQIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL------ | |||||||||||||||||||
| 6 | 3emlA | 0.23 | 0.24 | 0.89 | 3.83 | Download | ---------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVV-----------PMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLR | |||||||||||||||||||
| 7 | 4iaq | 0.22 | 0.24 | 0.85 | 1.71 | Download | -------------------SLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPF-FWRQASE----------CVVN----------TDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIAAARERKATKTLGIILGAFIVCWLPFFIISLVMPIH--LAIF-DFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRF---- | |||||||||||||||||||
| 8 | 2z73A | 0.15 | 0.21 | 0.95 | 2.67 | Download | W--YNPSIVVHPHWREFDQPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFG------WGAYTLEGVLC---------NSRDSTTRSNILCMFILGFFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTC | |||||||||||||||||||
| 9 | 3emlA | 0.23 | 0.24 | 0.89 | 4.93 | Download | -------------IM-------GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHI-INCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLR | |||||||||||||||||||
| 10 | 2ydoA | 0.19 | 0.25 | 0.92 | 5.62 | Download | -----------------S------SVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFAINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVYF | |||||||||||||||||||
| ||||||||||||||||||||||||||
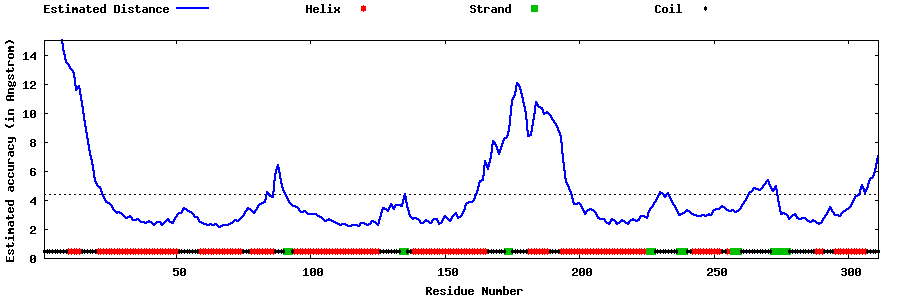
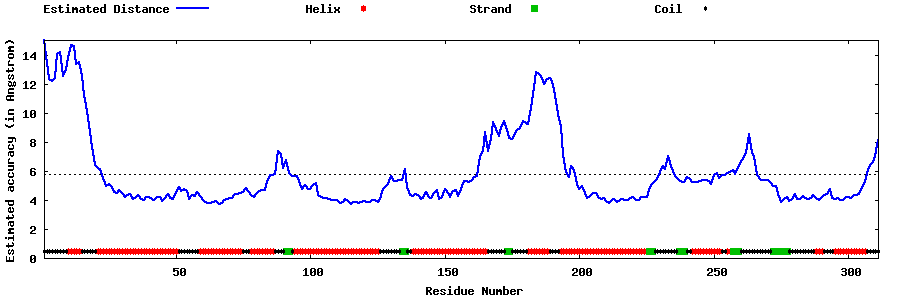
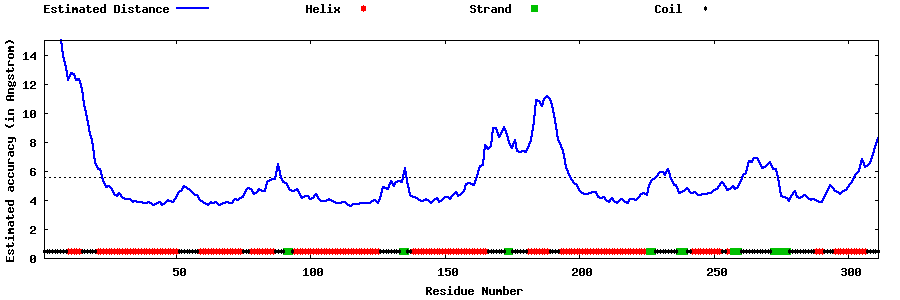
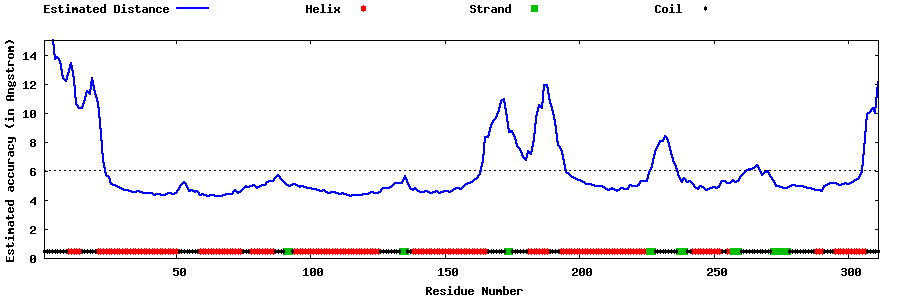
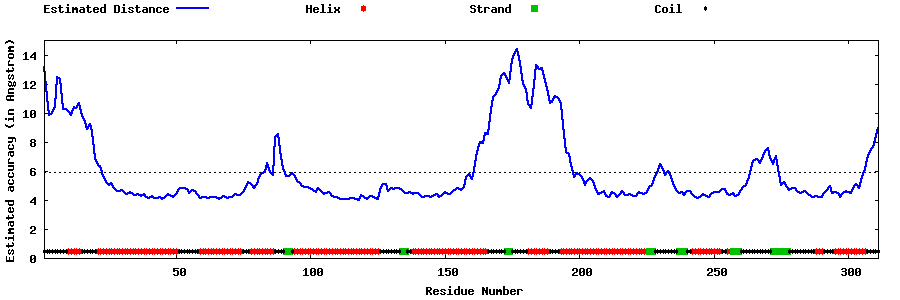
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||