

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| METANYTKVTEFVLTGLSQTREVQLVLFVIFLSFYLFILPGNILIICTIRLDPHLTSPMYFLLANLALLDIWYSSITAPKMLIDFFVERKIISFGGCIAQLFFLHFVGASEMFLLTVMAYDRYAAICRPLHYATIMNRRLCCILVALSWMGGFIHSIIQVALIVRLPFCGPNELDSYFCDITQVVRIACANTFPEELVMICSSGLISVVCFIALLMSYAFLLALLKKHSGSGENTNRAMSTCYSHITIVVLMFGPSIYIYARPFDSFSLDKVVSVFHTVIFPLLNPIIYTLRNKEVKAAMRKVVTKYILCEEK | |
| CCCCCCCSSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHCSSSSSCCCCCCCCCHHHHHHHHHHHHHHCCCHHHCCCCHHHHHHHHHHHHHHCCCCCC | |
| 9987787124777826999821589999999999999999788745523317999997899988699986633223729999987449967848999999999999999999999999866288757220262216875999999999999999999999995308998988377863281887888303405445225556769999999999999999999960116368877318878887999264663053258868899996313115777532440103743356299999999999752664679 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| METANYTKVTEFVLTGLSQTREVQLVLFVIFLSFYLFILPGNILIICTIRLDPHLTSPMYFLLANLALLDIWYSSITAPKMLIDFFVERKIISFGGCIAQLFFLHFVGASEMFLLTVMAYDRYAAICRPLHYATIMNRRLCCILVALSWMGGFIHSIIQVALIVRLPFCGPNELDSYFCDITQVVRIACANTFPEELVMICSSGLISVVCFIALLMSYAFLLALLKKHSGSGENTNRAMSTCYSHITIVVLMFGPSIYIYARPFDSFSLDKVVSVFHTVIFPLLNPIIYTLRNKEVKAAMRKVVTKYILCEEK | |
| 8566242200000000004244001000131333233233313300000100330300001002300120011000000200000035321000500000001103311310200020031000000221101000133100000020131022013001300230300001220000002210030000202301030022013102310331331121002001320463343110000030021003023210100000033313301000031033103311300002054023003301333032478 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCSSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHSSSSSCCCCCCCCHHHHHHHHHHHHHHHHCCCCHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCCCSSCCCHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHHHHHHHCSSSSSCCCCCCCCCHHHHHHHHHHHHHHCCCHHHCCCCHHHHHHHHHHHHHHCCCCCC METANYTKVTEFVLTGLSQTREVQLVLFVIFLSFYLFILPGNILIICTIRLDPHLTSPMYFLLANLALLDIWYSSITAPKMLIDFFVERKIISFGGCIAQLFFLHFVGASEMFLLTVMAYDRYAAICRPLHYATIMNRRLCCILVALSWMGGFIHSIIQVALIVRLPFCGPNELDSYFCDITQVVRIACANTFPEELVMICSSGLISVVCFIALLMSYAFLLALLKKHSGSGENTNRAMSTCYSHITIVVLMFGPSIYIYARPFDSFSLDKVVSVFHTVIFPLLNPIIYTLRNKEVKAAMRKVVTKYILCEEK | |||||||||||||||||||||||||
| 1 | 3emlA | 0.18 | 0.23 | 0.88 | 3.32 | Download | --------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVPMNYMVY-----------FNFFACVLVPLLLMLGVYLRIFLAARQLVHAAK--SLAIIVGLFALCWLPLHII-NCFTFFCPDCSHAWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ-- | |||||||||||||||||||
| 2 | 5tgzA | 0.21 | 0.28 | 0.89 | 2.26 | Download | -RGENFMDIECF----MVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRKD-SRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL--------GWNCEKL---------QSVCSDIFHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQRMDIELAKTLVLILVVLIICWGPLLAIMVKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSM---------- | |||||||||||||||||||
| 3 | 5tgzA | 0.20 | 0.28 | 0.88 | 2.04 | Download | GRGENFMD---------IENPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLCEKLQSVC-------------------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQRMDIELAKTLVLILVVLIICWGPLLAIMVGKMNLIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 4 | 4djh | 0.17 | 0.19 | 0.89 | 1.54 | Download | -------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLDRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSLSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------- | |||||||||||||||||||
| 5 | 4yay | 0.15 | 0.24 | 0.90 | 1.22 | Download | LKTTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVFF-------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK-----KN--DDIFKIIMAIVLFFFFSWIPHQIFTFLDVLIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL--------- | |||||||||||||||||||
| 6 | 5tgzA | 0.21 | 0.28 | 0.90 | 3.49 | Download | GRGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAV--------LPLLGWNCEK--------LQSVCSDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQAMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 7 | 4iaq | 0.19 | 0.22 | 0.85 | 1.73 | Download | --------------------LPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASECVVNT--------------------DHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADLARERKATKTLGIILGAFIVCWLPFFIISLVMPIHLAIF-DFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK------ | |||||||||||||||||||
| 8 | 2z73A | 0.16 | 0.22 | 0.96 | 3.12 | Download | -ETWNPSIVVHPHWREFDQVPAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGFPLMTISCFLKKIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGTLEGVLC------NCSFDY-----ISRDSTTRSNILCMFILGF--FGPILIIFFCYFNIVMMAKRLNAKEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPWVLTCCQF | |||||||||||||||||||
| 9 | 5tgzA | 0.20 | 0.28 | 0.90 | 4.49 | Download | GRGENFMDIECFMVL----NPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVF-HRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG---------NCEK---------LQSVCSDIFHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQRMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF--------- | |||||||||||||||||||
| 10 | 2ydoA | 0.19 | 0.21 | 0.92 | 5.32 | Download | -----------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQM-ESTLQKEVHAAKSLAIIVGLFALCCFTFFCPDCSHAPLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQE | |||||||||||||||||||
| ||||||||||||||||||||||||||
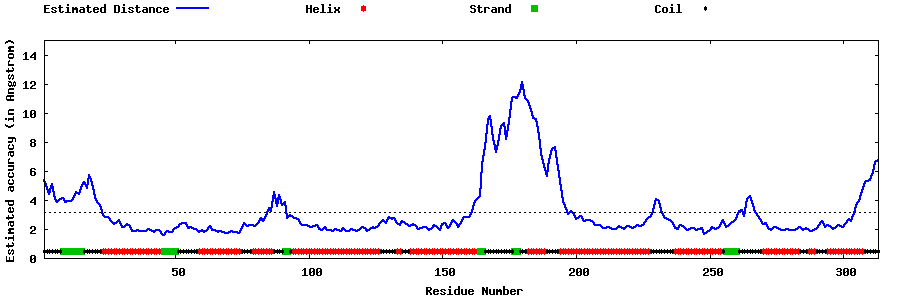
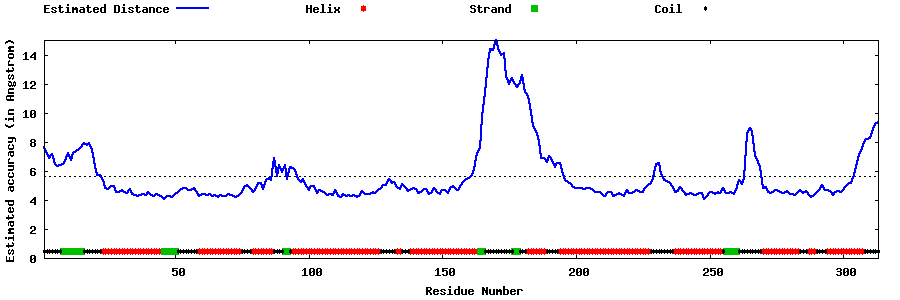
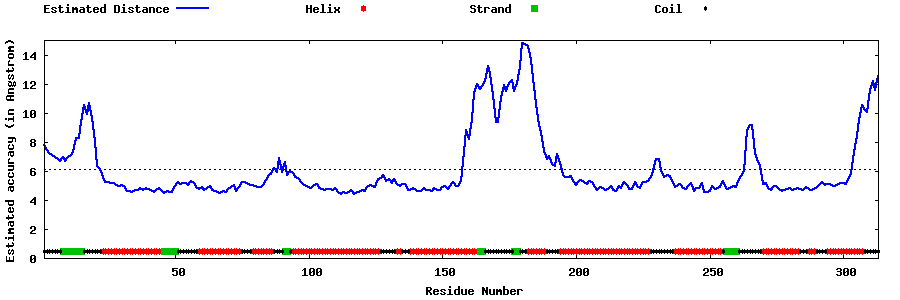
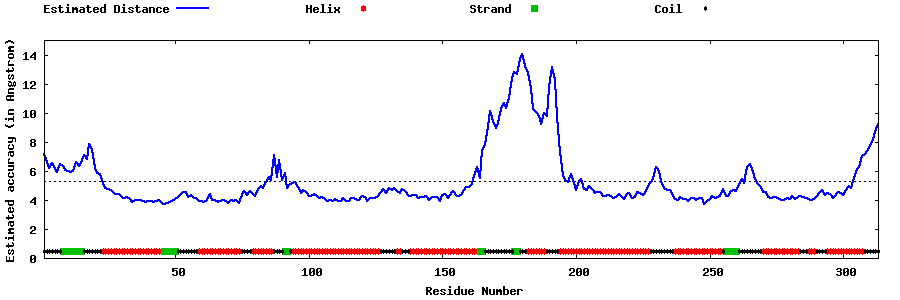
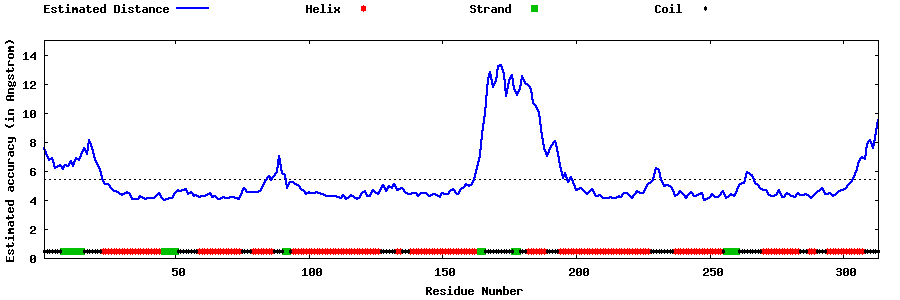
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||