

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MENYNQTSTDFILLGLFPQSRIGLFVFTLIFLIFLMALIGNLSMILLIFLDIHLHTPMYFLLSQLSLIDLNYISTIVPKMVYDFLYGNKSISFTGCGIQSFFFLTLAVAEGLLLTSMAYDRYVAICFPLHYPIRISKRVCVMMITGSWMISSINSCAHTVYALCIPYCKSRAINHFFCDVPAMLTLACTDTWVYESTVFLSSTIFLVLPFTGIACSYGRVLLAVYRMHSAEGRKKAYSTCSTHLTVVSFYYAPFAYTYVRPRSLRSPTEDKILAVFYTILTPMLNPIIYSLRNKEVMGALTQVIQKIFSVKM | |
| CCCCCCCSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCSSSSSHHCHHHCCCCHHHCCCCHHHHHHHHHHHHHHHHCCC | |
| 999984155788856999934388999999999999999899999999748875574999987799988744502579999986159968868999999999999999999999999865326206554488225887999999999999998999999999140789999778741482888888525850777789999999999979999999999999980477604364076526879998999987576315388999998779599872416243123654314659999999999987612059 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MENYNQTSTDFILLGLFPQSRIGLFVFTLIFLIFLMALIGNLSMILLIFLDIHLHTPMYFLLSQLSLIDLNYISTIVPKMVYDFLYGNKSISFTGCGIQSFFFLTLAVAEGLLLTSMAYDRYVAICFPLHYPIRISKRVCVMMITGSWMISSINSCAHTVYALCIPYCKSRAINHFFCDVPAMLTLACTDTWVYESTVFLSSTIFLVLPFTGIACSYGRVLLAVYRMHSAEGRKKAYSTCSTHLTVVSFYYAPFAYTYVRPRSLRSPTEDKILAVFYTILTPMLNPIIYSLRNKEVMGALTQVIQKIFSVKM | |
| 865522220100000004336120000130333233133323200200100030000001102000000010100100200000025432000200220110011001100200020020000000200200030033000000110021002002000000000201432301000013210030001003210210133123323312310331232001001103146123101000000200011132010000010324323733100001103310331030000104302400230033223144 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCSSSSSSSCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHCCHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCSSSSSHHCHHHCCCCHHHCCCCHHHHHHHHHHHHHHHHCCC MENYNQTSTDFILLGLFPQSRIGLFVFTLIFLIFLMALIGNLSMILLIFLDIHLHTPMYFLLSQLSLIDLNYISTIVPKMVYDFLYGNKSISFTGCGIQSFFFLTLAVAEGLLLTSMAYDRYVAICFPLHYPIRISKRVCVMMITGSWMISSINSCAHTVYALCIPYCKSRAINHFFCDVPAMLTLACTDTWVYESTVFLSSTIFLVLPFTGIACSYGRVLLAVYRMHSAEGRKKAYSTCSTHLTVVSFYYAPFAYTYVRPRSLRSPTEDKILAVFYTILTPMLNPIIYSLRNKEVMGALTQVIQKIFSVKM | |||||||||||||||||||||||||
| 1 | 3emlA | 0.18 | 0.20 | 0.89 | 3.51 | Download | -------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAK-SLAIIVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.22 | 0.83 | 2.12 | Download | ------------------NPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL------------------------PLLGWNCEKLQSVDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAMLCLLNSTVNPIIYALRSKDLRHAFRSM--------- | |||||||||||||||||||
| 3 | 5tgzA | 0.18 | 0.22 | 0.89 | 2.03 | Download | --GGRGENFMDIECFMVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRKD-SRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLIFPHID------------------KTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF-------- | |||||||||||||||||||
| 4 | 4djh | 0.15 | 0.19 | 0.90 | 1.53 | Download | ------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLGGTKVR---EDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP------ | |||||||||||||||||||
| 5 | 4yay | 0.18 | 0.20 | 0.91 | 1.25 | Download | KTTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVF-F------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLQLGIDIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL-------- | |||||||||||||||||||
| 6 | 3emlA | 0.19 | 0.20 | 0.89 | 3.74 | Download | --------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 7 | 4iaq | 0.20 | 0.23 | 0.84 | 1.70 | Download | --------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFF-WRQASECVVN--------------------TDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH--L-AIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRF------ | |||||||||||||||||||
| 8 | 4ea3A | 0.16 | 0.21 | 0.80 | 2.56 | Download | -------------------PGLKVTIVGLYLAVCVGGLLGNCLVMYVILRHTKMKTATNIYIFNLALADTLVLL-TLPFQGTDILLGFWPFGNALCKTVIAIDYYNMFTSTFTLTAMSVDRYVAICHPTSSKAQAVNVAIWALASVVGVPVAI--------MGSAQVEIPTPQDYWGPVFAICIFLFSF-----------------IVPVLVISVCYSLMIRRLRGVRLLSGSVAVFVGCWTPVQVFVLAQGLG-----VQPSSETAVAILRFCTALGYVNSCLNPILYAFLDENFKACFR----------- | |||||||||||||||||||
| 9 | 3emlA | 0.18 | 0.20 | 0.89 | 4.77 | Download | -------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKS-LAIIVGLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ- | |||||||||||||||||||
| 10 | 2ydoA | 0.17 | 0.22 | 0.92 | 5.67 | Download | ----------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWLPFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQQ | |||||||||||||||||||
| ||||||||||||||||||||||||||
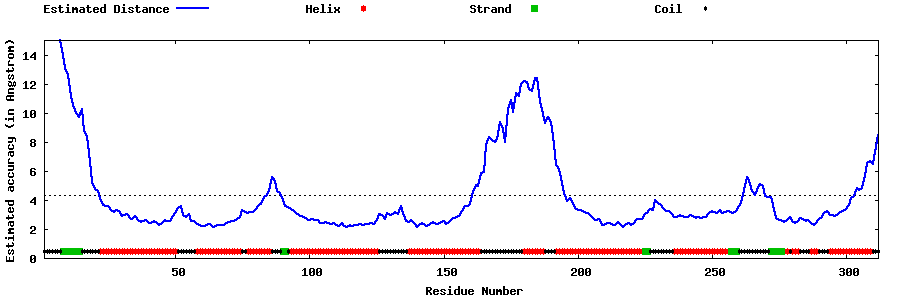
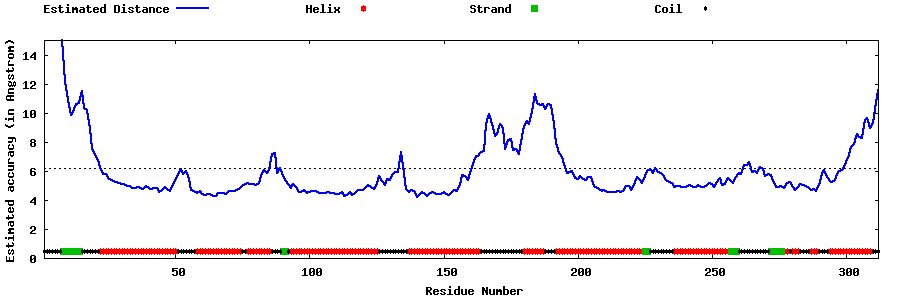
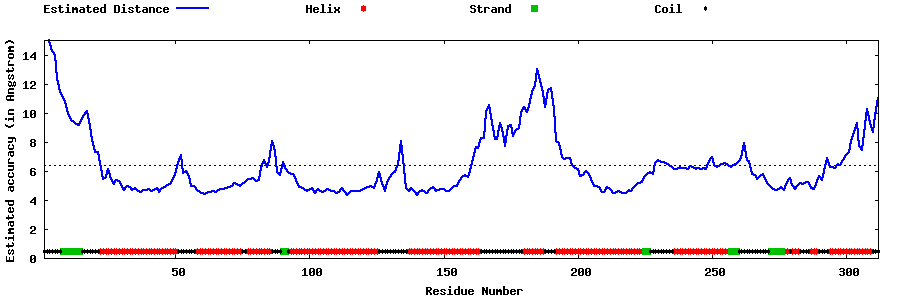
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||