

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTNLNASQANHRNFILTGIPGTPDKNPWLAFPLGFLYTLTLLGNGTILAVIKVEPSLHEPTYYFLSILALTDVSLSMSTLPSMLSIYWFNAPQIVFDACIMQMFFIHVFGIVESGVLVSMAFDRFVAIRNPLHYVSILTHDVIRKTGIAVLTRAVCVVFPVPFLIKCLPFCHSNVLSHSYCLHQNMMRLACASTRINSLYGLIVVIFTLGLDVLLTLLSYVLTLKTVLGIVSRGERLKTLSTCLSHMSTVLLFYVPFMGAASMIHRFWEHLSPVVHMVMADIYLLLPPVLNPIVYSVKTKQI | |
| CCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCCCCC | |
| 99989998834101268998986789999999999999999999999998857887533189999999999999998449999999956998688889999999999999999999999982417741666456311888899999999999999999889999943999998946874431065688824784376699999999999998999999999999998078897788898660699999999999989988761046888899478999999996378553682442037769 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MTNLNASQANHRNFILTGIPGTPDKNPWLAFPLGFLYTLTLLGNGTILAVIKVEPSLHEPTYYFLSILALTDVSLSMSTLPSMLSIYWFNAPQIVFDACIMQMFFIHVFGIVESGVLVSMAFDRFVAIRNPLHYVSILTHDVIRKTGIAVLTRAVCVVFPVPFLIKCLPFCHSNVLSHSYCLHQNMMRLACASTRINSLYGLIVVIFTLGLDVLLTLLSYVLTLKTVLGIVSRGERLKTLSTCLSHMSTVLLFYVPFMGAASMIHRFWEHLSPVVHMVMADIYLLLPPVLNPIVYSVKTKQI | |
| 44442434231310200000121320200011022022203321210000020141002000200010030020003002020000000214603140000001301310331110011002000000021010000013410010000013212011202101021031046200000001021001000230300111012003313333231032012200200030124601320120010000000002123133000001104310110001103310130033003000132358 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHSSCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCCCCC MTNLNASQANHRNFILTGIPGTPDKNPWLAFPLGFLYTLTLLGNGTILAVIKVEPSLHEPTYYFLSILALTDVSLSMSTLPSMLSIYWFNAPQIVFDACIMQMFFIHVFGIVESGVLVSMAFDRFVAIRNPLHYVSILTHDVIRKTGIAVLTRAVCVVFPVPFLIKCLPFCHSNVLSHSYCLHQNMMRLACASTRINSLYGLIVVIFTLGLDVLLTLLSYVLTLKTVLGIVSRGERLKTLSTCLSHMSTVLLFYVPFMGAASMIHRFWEHLSPVVHMVMADIYLLLPPVLNPIVYSVKTKQI | |||||||||||||||||||||||||
| 1 | 3emlA | 0.16 | 0.23 | 0.87 | 2.37 | Download | ---------------------IM-GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACL-----------FEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC----PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREF | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.23 | 0.87 | 2.00 | Download | -----------------------SQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-------GWNCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDL | |||||||||||||||||||
| 3 | 5tgzA | 0.18 | 0.23 | 0.91 | 1.87 | Download | -----GGRGENFMDIECFMVLNPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLIFPHID------------------KTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFAFCSMLCLLNSTVNPIIYALRSKDL | |||||||||||||||||||
| 4 | 3uon | 0.16 | 0.23 | 0.88 | 1.56 | Download | ----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIV--GVRTVEDGECYIQF---------FSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSRINISREKKVTRTILAILLAFIITWAPYN-VMVLINTFCAPIPNTV-WTIGYWLCYINSTINPACYALCNATF | |||||||||||||||||||
| 5 | 3uonA | 0.17 | 0.20 | 0.87 | 1.14 | Download | ----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFI--VGVRTVEDGECYIQFF---------SNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKS----RREKKVTRTILAILLAFIITWAPYNV-MVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATF | |||||||||||||||||||
| 6 | 3emlA | 0.16 | 0.23 | 0.87 | 2.64 | Download | -----------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVV-----------PMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF----TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREF | |||||||||||||||||||
| 7 | 4iaq | 0.19 | 0.23 | 0.62 | 1.69 | Download | ----------------------LPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFF-WRQASECVVN--------------------TDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSR------------------------------------------------------------------------ | |||||||||||||||||||
| 8 | 4buoA | 0.14 | 0.18 | 0.91 | 2.45 | Download | ----NSDL---------DVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLARKKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHHPWAFAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPG------------GVDTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQRGVLVLRAVVIAFVVCWLPYHVRRLMFCYI---SDEQWTYHYFYMLTNALVYVSAAINPILYNLVSANF | |||||||||||||||||||
| 9 | 5tgzA | 0.19 | 0.23 | 0.91 | 3.07 | Download | -GGRGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG--------NCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKA-APDQARMDIELAKTLVLILVVLIICWGPLLAIMVYD--FGKMNKIKTVFAFCSMLCLLNSTVNPIIYALRSKDL | |||||||||||||||||||
| 10 | 4gpoA | 0.17 | 0.21 | 0.90 | 4.82 | Download | ---------------------LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPG--------CCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQVMLMREHKALKTLGIIMGVFTLCWLPFFLVNI-VNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDF | |||||||||||||||||||
| ||||||||||||||||||||||||||
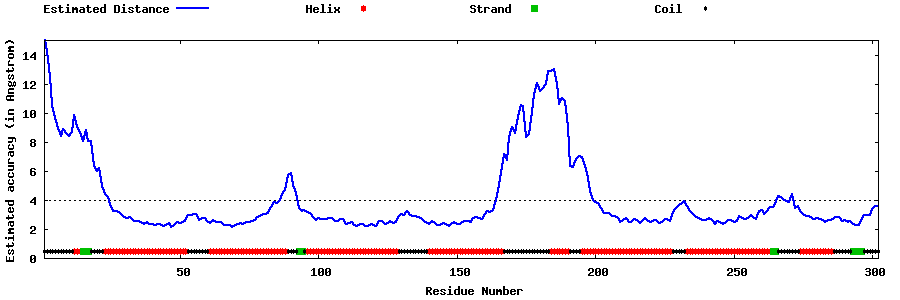
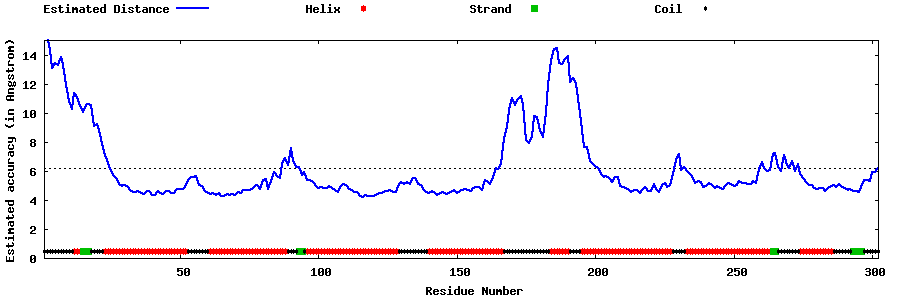
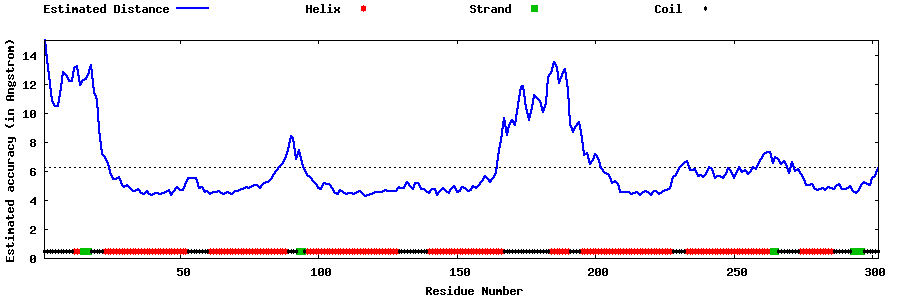
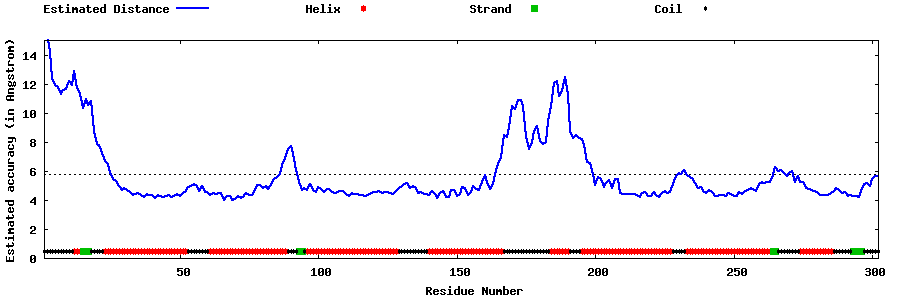
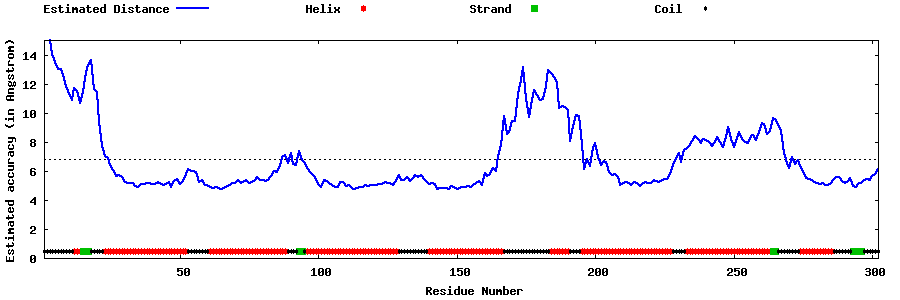
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||