

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MSPDTFLKGFAEFFLMGFSNSWDIQIVHAALFFLVYLAAVIGNLLIIILTTLDVHLQTPMYFFLRNLSFLDFCYISVTIPKSIVSSLTHDTSISFFGCALQAFFFMDLATTEVAILTVMSYDRYMAICRPLHYEVIINQGVCLRMMAMSWLSGVICGFMHVIATFSLPFCGRNRIRQFFCNIPQLLSLLDPKVITIEIGVMVFGTSLVIISFVVITLSYMYIFSVIMRIPSKEGRSKTFSTCIPHLVVVTLFMISGSIAYVKPISNSPPVLDVFLSAFYTVVPPTLNPVIYSLRNRDMKAALRRQCGP | |
| CCCCCCCSSCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHCCHHHHHHHHCCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHCCHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHCC | |
| 99988723203330015899850799999999999999999889999999708875675888987899997757712589999985269976868999999999999999999999999865078626554577244885999999999999999999999999144899949378864586898888414634999999999999999989999999999999971386505565332070848997999985477358179999988778189874308021102454414539999999997477 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MSPDTFLKGFAEFFLMGFSNSWDIQIVHAALFFLVYLAAVIGNLLIIILTTLDVHLQTPMYFFLRNLSFLDFCYISVTIPKSIVSSLTHDTSISFFGCALQAFFFMDLATTEVAILTVMSYDRYMAICRPLHYEVIINQGVCLRMMAMSWLSGVICGFMHVIATFSLPFCGRNRIRQFFCNIPQLLSLLDPKVITIEIGVMVFGTSLVIISFVVITLSYMYIFSVIMRIPSKEGRSKTFSTCIPHLVVVTLFMISGSIAYVKPISNSPPVLDVFLSAFYTVVPPTLNPVIYSLRNRDMKAALRRQCGP | |
| 66776130201100000004334000110230332133133333200200100430000001002000000010100100200000016432000100120110001002100000010010000000200200020033000000000021013103100010000200361402000012220030001203110111123113313312320330233001000103166323100000110200000133002100010316334424100002103331330030000204201400330258 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCSSCHHHHSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHCCHHHHHHHHCCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHCCHHHHHHHHHHHHHHHHHSSSCCCCCCCCCCCCSSSSSCCCCCCCCCCHHHCCCCHHHHHHHHHHHCC MSPDTFLKGFAEFFLMGFSNSWDIQIVHAALFFLVYLAAVIGNLLIIILTTLDVHLQTPMYFFLRNLSFLDFCYISVTIPKSIVSSLTHDTSISFFGCALQAFFFMDLATTEVAILTVMSYDRYMAICRPLHYEVIINQGVCLRMMAMSWLSGVICGFMHVIATFSLPFCGRNRIRQFFCNIPQLLSLLDPKVITIEIGVMVFGTSLVIISFVVITLSYMYIFSVIMRIPSKEGRSKTFSTCIPHLVVVTLFMISGSIAYVKPISNSPPVLDVFLSAFYTVVPPTLNPVIYSLRNRDMKAALRRQCGP | |||||||||||||||||||||||||
| 1 | 3emlA | 0.20 | 0.20 | 0.88 | 3.59 | Download | ---------------------IMGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS---GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINC-FTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 2 | 5tgzA | 0.20 | 0.25 | 0.92 | 2.15 | Download | -GRGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVL--------PLLGWNCEKL---------QSVCSDIFPHDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFSMLCLLNSTVNPIIYALRSKDLRHAFRSMF-- | |||||||||||||||||||
| 3 | 5tgzA | 0.19 | 0.25 | 0.90 | 2.23 | Download | -----------GGRGENFMDIPSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLQSV-------------CSDIFPHIKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF-- | |||||||||||||||||||
| 4 | 4djh | 0.16 | 0.23 | 0.91 | 1.55 | Download | --------------------SPAIPVIITAVYSVVFVVGLVGNSLVMFVIIRYTKMKTATNIYIFNLALADALVTT-TMPFQSTVYLMNSWPFGDVLCKIVLSIDYYNMFTSIFTLTMMSVDRYIAVCHPVKALDFRTPLKAKIINICIWLLSSSVGISAIVLG-GTKV--REDVDVIECSLQFPDD---DYSWWDLFMKICVFIFAFVIPVLIIIVCYTLMILRLKSVRLLRNLRRITRLVLVVVAVFVVCWTPIHIFILVEALGSAALSSYYFCIALGYTNSSLNPILYAFLDENFKRCFRDFCFP | |||||||||||||||||||
| 5 | 4yay | 0.16 | 0.21 | 0.93 | 1.23 | Download | AEKTTRNAYIQKYLILNSSDCNYIFVMIPTLYSIIFVVGIFGNSLVVIVIYFYMKLKTVASVFLLNLALADLCFLLTLPLWAVYTAMEYRWPFGNYLCKIASASVSFNLYASVFLLTCLSIDRYLAIVHPMKSRLRRTMLVAKVTCIIIWLLAGLASLPAIIHRNVF-F------IITVCAFHYE--------TLPIGLGLTKNILGFLFPFLIILTSYTLIWKALK------KNDDIFKIIMAIVLFFFFSWIPHQIFTFLQLGIDIVDTAMPITICIAYFNNCLNPLFYGFLGKKFKRYFLQLL-- | |||||||||||||||||||
| 6 | 3emlA | 0.20 | 0.20 | 0.88 | 3.79 | Download | ----------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAIT--ISTGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCF-TFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 7 | 3d4s | 0.19 | 0.23 | 0.64 | 1.72 | Download | ------------------------VVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIWTLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMHWYRA--TH--QEAINCYAEE----TCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKR-------------------------------------------------------------------------------- | |||||||||||||||||||
| 8 | 2z73A | 0.17 | 0.22 | 0.94 | 2.56 | Download | ---ETNPSIVVHPHWREFDQPDAVYYSLGIFIGICGIIGCGGNGIVIYLFTKTKSLQTPANMFIINLAFSDFTFSLVNGPLMTISCFLKKWIFGFAACKVYGFIGGIFGFMSIMTMAMISIDRYNVIGRPMAASKKMSHRRAFIMIIFVWLWSVLWAIGPIFGTLEGVLCNDYI----------------SRDSTTRSNILCMFILGFFGPILIIFFCYFNIVMAQAGANAEMRLAKISIVIVSQFLLSWSPYAVVALLAQFGPLEWVTPYAAQLPVMFAKASAIHNPMIYSVSHPKFREAISQTFPW | |||||||||||||||||||
| 9 | 3emlA | 0.19 | 0.20 | 0.89 | 4.76 | Download | -------------IM--------GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIS--TGFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLNCGQSQGCGEGQVACLFEDVVP-----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIV-GLFALCWLPLHIINCFTFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| 10 | 2ydoA | 0.17 | 0.20 | 0.92 | 5.52 | Download | ------------------------SSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSAAAADILVGVLAIPFAIA--ISTGFCAACHGCLFIACFVLVLTASSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGQPKEGKAHSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLKQMESTLQKEVHAAKSLAIIVGLFALCWLPFFCPDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRS | |||||||||||||||||||
| ||||||||||||||||||||||||||
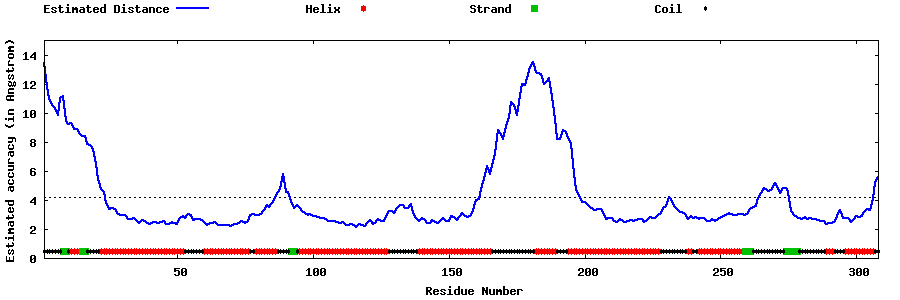
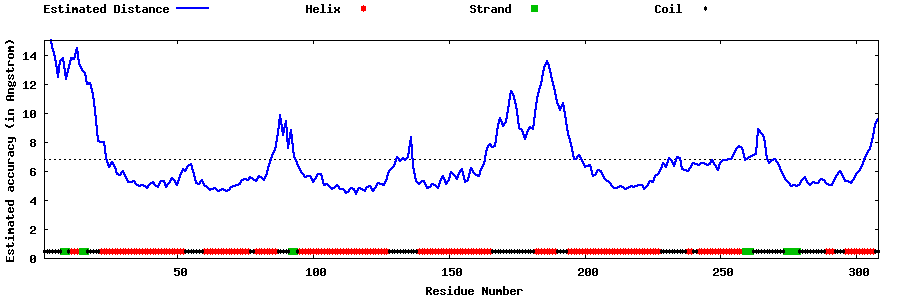
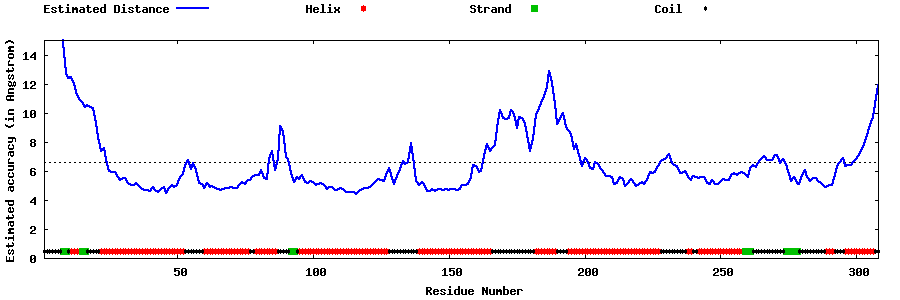
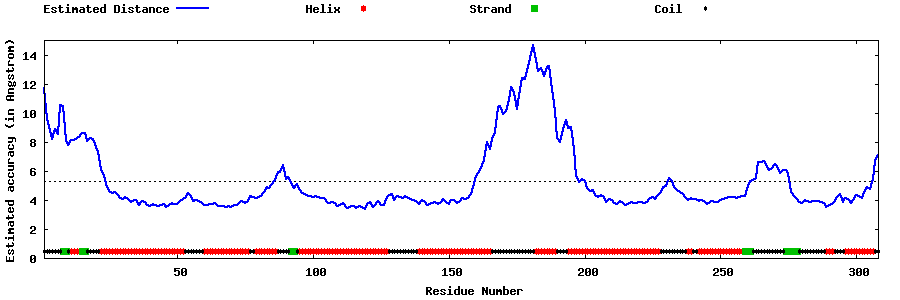
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||