

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MPIANDTQFHTSSFLLLGIPGLEDVHIWIGFPFFSVYLIALLGNAAIFFVIQTEQSLHEPMYYCLAMLDSIDLSLSTATIPKMLGIFWFNIKEISFGGYLSQMFFIHFFTVMESIVLVAMAFDRYIAICKPLWYTMILTSKIISLIAGIAVLRSLYMVIPLVFLLLRLPFCGHRIIPHTYCEHMGIARLACASIKVNIMFGLGSISLLLLDVLLIILSHIRILYAVFCLPSWEARLKALNTCGSHIGVILAFSTPAFFSFFTHCFGHDIPQYIHIFLANLYVVVPPTLNPVIYGVRTKHIRETVLRIFFKTDH | |
| CCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHCCCC | |
| 9998999874521404799898668899999999999999999999999885788754308999999999999999967999999995699868888899999999999999999999998342775166654631278899999999999999999998999996389999994587121006567871578278699999999999998999999999999998268887788898650599999999999989988423115798981799999999973886646801116579999999998510579 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MPIANDTQFHTSSFLLLGIPGLEDVHIWIGFPFFSVYLIALLGNAAIFFVIQTEQSLHEPMYYCLAMLDSIDLSLSTATIPKMLGIFWFNIKEISFGGYLSQMFFIHFFTVMESIVLVAMAFDRYIAICKPLWYTMILTSKIISLIAGIAVLRSLYMVIPLVFLLLRLPFCGHRIIPHTYCEHMGIARLACASIKVNIMFGLGSISLLLLDVLLIILSHIRILYAVFCLPSWEARLKALNTCGSHIGVILAFSTPAFFSFFTHCFGHDIPQYIHIFLANLYVVVPPTLNPVIYGVRTKHIRETVLRIFFKTDH | |
| 4444242423121020000012132020001102201220332121000002015100100020001001000000200002000000020340304000000120031023010001100300000002101000000341001000000321201120210102103204520000000102100100023030011100201313331132032011100200010024601320120010010002003231331010011033102100010022100300330030000203400410030024378 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCHHHSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCHHHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHCCCC MPIANDTQFHTSSFLLLGIPGLEDVHIWIGFPFFSVYLIALLGNAAIFFVIQTEQSLHEPMYYCLAMLDSIDLSLSTATIPKMLGIFWFNIKEISFGGYLSQMFFIHFFTVMESIVLVAMAFDRYIAICKPLWYTMILTSKIISLIAGIAVLRSLYMVIPLVFLLLRLPFCGHRIIPHTYCEHMGIARLACASIKVNIMFGLGSISLLLLDVLLIILSHIRILYAVFCLPSWEARLKALNTCGSHIGVILAFSTPAFFSFFTHCFGHDIPQYIHIFLANLYVVVPPTLNPVIYGVRTKHIRETVLRIFFKTDH | |||||||||||||||||||||||||
| 1 | 3emlA | 0.20 | 0.24 | 0.88 | 2.48 | Download | ---------------------IM-GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDV----------VPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHRQ | |||||||||||||||||||
| 2 | 5tgzA | 0.18 | 0.23 | 0.85 | 2.13 | Download | -----------------------SQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSV-----------------CSDIFIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHAPDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 3 | 5tgzA | 0.18 | 0.23 | 0.88 | 1.93 | Download | FMDIECFMVLNP-----------SQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFH-VFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG----WNCEKLIFPHID-----------------KTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGK-MNIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 4 | 3uon | 0.17 | 0.20 | 0.88 | 1.56 | Download | ----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFIV--GVRTVEDGECYIQFF---------SNAAVTFGTAAAFYLPVIIMTVLYWHISRASKSRINISREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPIPNTV-WTIGYWLCYINSTINPACYALCNATFKKTFKHLLM---- | |||||||||||||||||||
| 5 | 3uonA | 0.17 | 0.20 | 0.87 | 1.12 | Download | ----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFI--VGVRTVEDGECYIQF---------FSNAAVTFGTIAAFYLPVIIMTVLYWHISRASKS----RREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM---- | |||||||||||||||||||
| 6 | 3emlA | 0.20 | 0.24 | 0.88 | 2.66 | Download | -----------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVVP----------MNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| 7 | 4iaq | 0.18 | 0.20 | 0.84 | 1.70 | Download | -----------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPFFW-RQASECVVNT-----------------D---HILYVYSTVGAFYFPTLLLIALYGRIYVEARSRIADARERKATKTLGIILGAFIVCWLPFFIISLVMPIH---L-AIFDF-FTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK-- | |||||||||||||||||||
| 8 | 4buoA | 0.17 | 0.21 | 0.92 | 2.81 | Download | ----NSD---------LDVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLARKKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHHPWAFAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPGGLVCTPIV------DIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQPGRVQALRRGVLVLRAVVIAFVVCWLPYHVRRLMFCYISDEQWTYHYFYMLTNALVYVSAAINPILYNLVSANFRQVFLSTL----- | |||||||||||||||||||
| 9 | 5tgzA | 0.19 | 0.23 | 0.90 | 3.25 | Download | GRGENFMDIEC--FMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG-------NCEKLQSV-----------CSDIHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKA-APDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 10 | 4gpoA | 0.18 | 0.21 | 0.88 | 4.74 | Download | ---------------------LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPG--------CCDFVTNRAYAIASSIIFYIPLLIMIFVALRVYREAKEQVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRL------ | |||||||||||||||||||
| ||||||||||||||||||||||||||
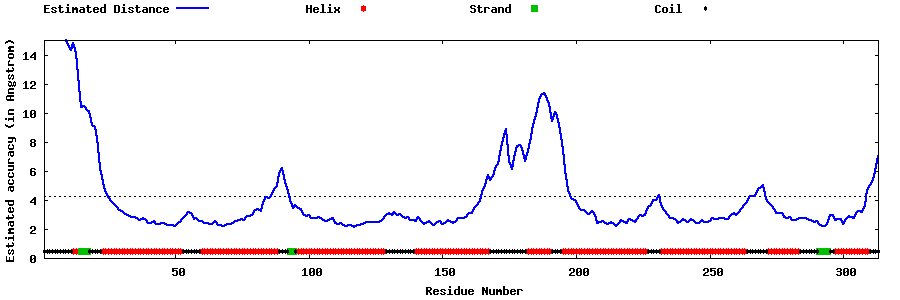
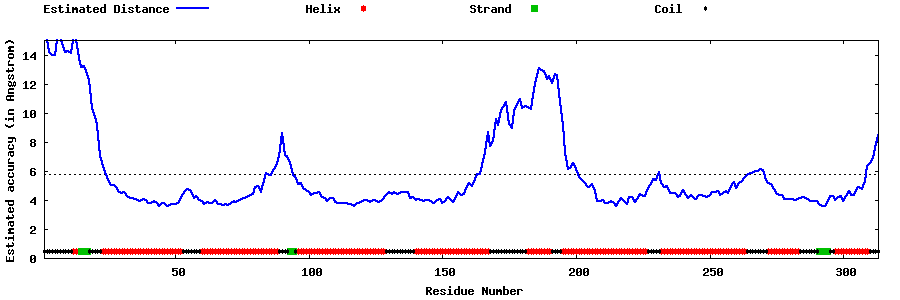
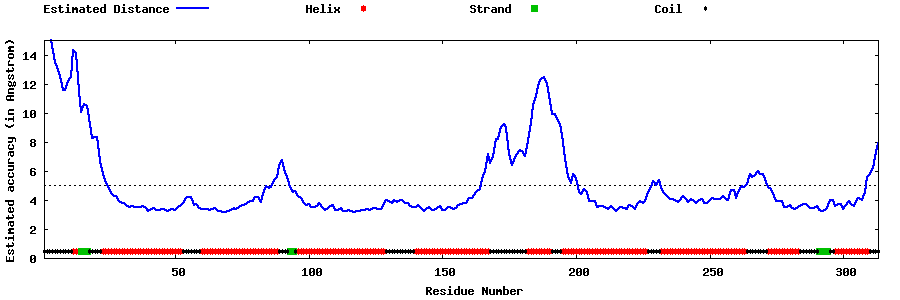
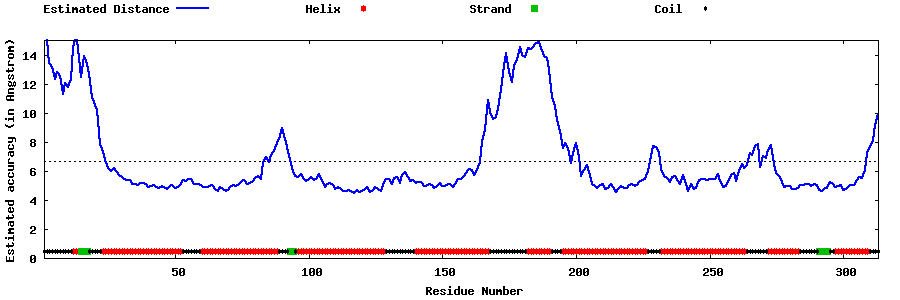
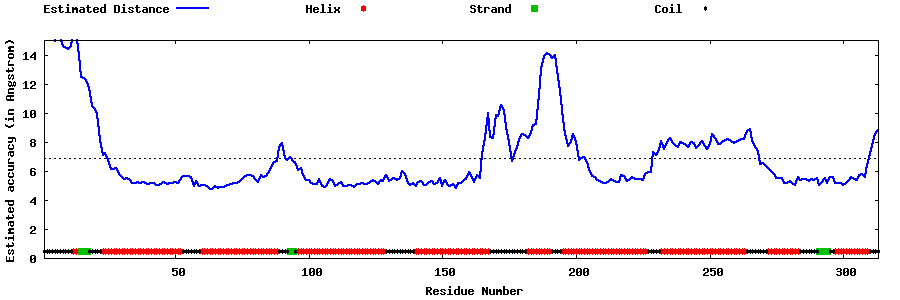
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||