

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MKISNNSLGFLPTTFILVGIPGLESEHLWISVPFSLIYIIIFLGNGIILHVIRTDIALHQPMYLFLAMLALAEVRVSASTLPTVLGIFLFGNTEISLEACLFPDVLHPFFIHDGASCAAGHVFGPLYSHLQPTELHSYPDTAQGLWHRSYYRTEKHYAHGSVAHSLMASALLWPQCPLTFLLSAPQSYLSCGNISVNNIYGIFIVTSTFGLDSLLIVISYGLILHTVLGIATGEGRKKALNTCGSHVCAVLAYYVPMIGLSIVHRLGHRVSPLLQAMMANAYLFFPPVVNPIVYSIKTKEIHGAIVRMLLEKRRRV | |
| CCCCCCCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHHCCCCC | |
| 9987799987564124779999854899999999999999999999999999668975432899999999999999985499999999569988878999999999999999999999999988499987733035123401736877657988999999999999999980587888322307678883478267679999999999999899999999999999818888758889867079999999999999999971044678788425478878885278765680010658999999999987645779 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MKISNNSLGFLPTTFILVGIPGLESEHLWISVPFSLIYIIIFLGNGIILHVIRTDIALHQPMYLFLAMLALAEVRVSASTLPTVLGIFLFGNTEISLEACLFPDVLHPFFIHDGASCAAGHVFGPLYSHLQPTELHSYPDTAQGLWHRSYYRTEKHYAHGSVAHSLMASALLWPQCPLTFLLSAPQSYLSCGNISVNNIYGIFIVTSTFGLDSLLIVISYGLILHTVLGIATGEGRKKALNTCGSHVCAVLAYYVPMIGLSIVHRLGHRVSPLLQAMMANAYLFFPPVVNPIVYSIKTKEIHGAIVRMLLEKRRRV | |
| 5433324232312202000001213202000110220122033212100000211410020002000000200100030020200000002144010300133210000000310110000000102331221010000000210100000224300010000003213012001000000002100100023030011101200331333323103201110020001002560132012001000000000321133101001003310210000002210130133003000020340041003002444665 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHHCCCCC MKISNNSLGFLPTTFILVGIPGLESEHLWISVPFSLIYIIIFLGNGIILHVIRTDIALHQPMYLFLAMLALAEVRVSASTLPTVLGIFLFGNTEISLEACLFPDVLHPFFIHDGASCAAGHVFGPLYSHLQPTELHSYPDTAQGLWHRSYYRTEKHYAHGSVAHSLMASALLWPQCPLTFLLSAPQSYLSCGNISVNNIYGIFIVTSTFGLDSLLIVISYGLILHTVLGIATGEGRKKALNTCGSHVCAVLAYYVPMIGLSIVHRLGHRVSPLLQAMMANAYLFFPPVVNPIVYSIKTKEIHGAIVRMLLEKRRRV | |||||||||||||||||||||||||
| 1 | 3emlA | 0.14 | 0.17 | 0.88 | 2.59 | Download | ----------------------IM-GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTR--AKGIIAICWVLSFAIGLTPMLG-----WNNCGQSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSH-RQ- | |||||||||||||||||||
| 2 | 5tgzA | 0.15 | 0.19 | 0.85 | 2.19 | Download | -----------------------PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRCPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPK------AVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARMELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF------- | |||||||||||||||||||
| 3 | 4z34A | 0.14 | 0.17 | 0.90 | 1.43 | Download | QCFYNESIAFFYNRSGKHLATEWNTVSKLVMGLGITVCIFIMLANLLVMVAIYVNRRFHFPIYYLMANLAAADFFAGLAYFYLMFNTGPNTRR---LTV--STWLLRQGLIDTSLTASVANLLAIAIERHITVF---------RMQLHTRMSNRRVVVVIVVIWTMAIVMGAIPSVWNCICD-----IENCSNMALYSDSYLVFWAIFNLVTFVVMVVLYAH------YLRNRDTMMSLLKTVVIVLGAFIICWTPGLVLLLLDVCCPQCDVLAYEKFFLLLAEFNSAMNPIIYSYRDKEMSATFRQILG------ | |||||||||||||||||||
| 4 | 4ib4 | 0.17 | 0.19 | 0.85 | 1.51 | Download | --------------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFAMWPLPLVLCPAWL----FLDVLFSTASIWHLVDRYIAIKKPIQYNSRATAFITVWLISIGI--------AIPVPIK----GIETNP---------NNITCVLTKEFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAAISNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCNQQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR-- | |||||||||||||||||||
| 5 | 3uonA | 0.12 | 0.15 | 0.86 | 1.13 | Download | -----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAG--MMIAAAWVLSFILWAPAILFWQFIVGRTVEDGECYIQF---------FSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKS----RREKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM------ | |||||||||||||||||||
| 6 | 3emlA | 0.14 | 0.17 | 0.89 | 2.87 | Download | ------------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVT--GTRAKGIIAICWVLSFAIGLTPMLG-----WNNCGQSQGCGEGQVACLFEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVLRQ | |||||||||||||||||||
| 7 | 4iaq | 0.15 | 0.20 | 0.83 | 1.70 | Download | ------------------------PWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSI--SISLPPF------FWR--QASEC----------VVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIAAARERKATKTLGIILGAFIVCWLPFFIISLVMPIH----LAIFDF-FTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK---- | |||||||||||||||||||
| 8 | 1u19A | 0.16 | 0.25 | 0.92 | 1.88 | Download | MNPFSNKTGVVRSPF---EAPQYPWQFSMLAAYMFLLIMLGFPINFLTLYVTVQHKKLRTPLNYILLNLAVADLFMVFGGFTTTLYTSLHGYFVFGPTGCNLEGFFATLGGEIALWSLVVLAIERYVVVCKPMSNFRF-------------GENHAIMGVAFTWVMALACAAPPLVGWSYYT-------PHEETN-NESFVIYMFVVHFIIPLIVIFFCYGQLVFTVESATTQKAEKEVTRMVIIMVIAFLICWLPVAFYIFTHQGSD--FGPIFMTIPAFFAKTSAVYNPVIYIMMNKQFRNCMVTTLCCGKNPL | |||||||||||||||||||
| 9 | 5tgzA | 0.14 | 0.19 | 0.91 | 3.08 | Download | --GGRGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVT------RPKAVVAFCLMWTIAIVIAVLPLLGWNCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKA-APDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF------- | |||||||||||||||||||
| 10 | 4gpoA | 0.17 | 0.19 | 0.87 | 4.21 | Download | ----------------------LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRA--RAKVIICTVWAISALVSFLPIMMHWWRDEDLKCYQDPG--------CCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRL-------- | |||||||||||||||||||
| ||||||||||||||||||||||||||
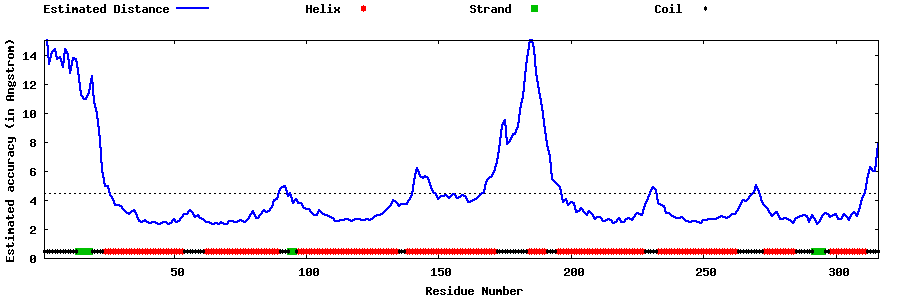
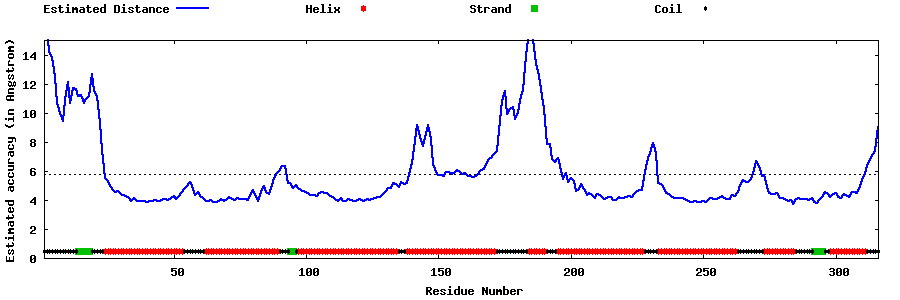
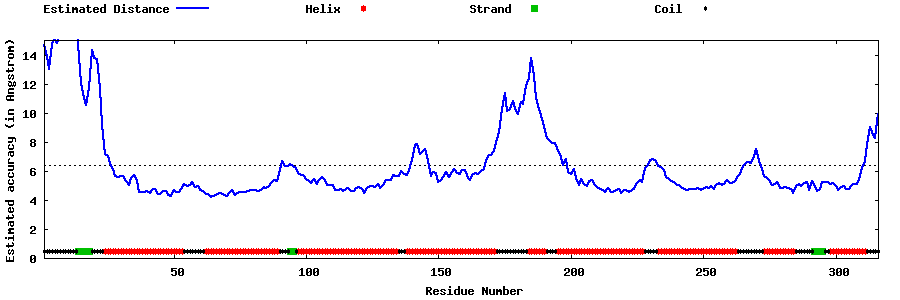
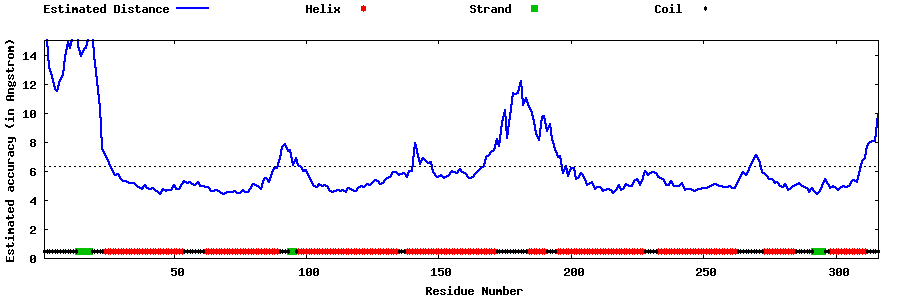
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||