

| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MLGLNGTPFQPATLQLTGIPGIQTGLTWVALIFCILYMISIVGNLSILTLVFWEPALHQPMYYFLSMLALNDLGVSFSTLPTVISTFCFNYNHVAFNACLVQMFFIHTFSFMESGILLAMSLDRFVAICYPLRYVTVLTHNRILAMGLGILTKSFTTLFPFPFVVKRLPFCKGNVLHHSYCLHPDLMKVACGDIHVNNIYGLLVIIFTYGMDSTFILLSYALILRAMLVIISQEQRLKALNTCMSHICAVLAFYVPIIAVSMIHRFWKSAPPVVHVMMSNVYLFVPPMLNPIIYSVKTKEIRKGILKFFHKSQA | |
| CCCCCCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHCCCC | |
| 99988997726602477899976788999999999999999999999999965787543089999999999998998679999999946998788878999999999999999999999980417862665457703868999999999999999999889999823899998967874542132567714795042299999999999999999999999999998078898898999761799999999999999998600347998884589999999974785536811105479999999998500479 |
| 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | |
| | | | | | | | | | | | | | | | | |
| MLGLNGTPFQPATLQLTGIPGIQTGLTWVALIFCILYMISIVGNLSILTLVFWEPALHQPMYYFLSMLALNDLGVSFSTLPTVISTFCFNYNHVAFNACLVQMFFIHTFSFMESGILLAMSLDRFVAICYPLRYVTVLTHNRILAMGLGILTKSFTTLFPFPFVVKRLPFCKGNVLHHSYCLHPDLMKVACGDIHVNNIYGLLVIIFTYGMDSTFILLSYALILRAMLVIISQEQRLKALNTCMSHICAVLAFYVPIIAVSMIHRFWKSAPPVVHVMMSNVYLFVPPMLNPIIYSVKTKEIRKGILKFFHKSQA | |
| 44432324231310200000121320200011022022203321210000020141002000200010020010003002020000000214603040000001201310230120011002000000021010000003410010000002212011202101021021046200000001021001000230300111012003313333231032011200200020014601320120010000000003211321010010133101100000022100300330030000203300410030026468 |
| Rank | PDB Hit | Iden1 | Iden2 | Cov. | Norm. Z-score | Download Align. | 20 40 60 80 100 120 140 160 180 200 220 240 260 280 300 | | | | | | | | | | | | | | | | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sec.Str Seq | CCCCCCCCCCCCSSSSSCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCSSCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCCCCCCCCCHHHHHHHCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHCCCCCCCCCHHHHHHHHHHHHCCCCCCCSSSSCCHHHHHHHHHHHHHCCCC MLGLNGTPFQPATLQLTGIPGIQTGLTWVALIFCILYMISIVGNLSILTLVFWEPALHQPMYYFLSMLALNDLGVSFSTLPTVISTFCFNYNHVAFNACLVQMFFIHTFSFMESGILLAMSLDRFVAICYPLRYVTVLTHNRILAMGLGILTKSFTTLFPFPFVVKRLPFCKGNVLHHSYCLHPDLMKVACGDIHVNNIYGLLVIIFTYGMDSTFILLSYALILRAMLVIISQEQRLKALNTCMSHICAVLAFYVPIIAVSMIHRFWKSAPPVVHVMMSNVYLFVPPMLNPIIYSVKTKEIRKGILKFFHKSQA | |||||||||||||||||||||||||
| 1 | 3emlA | 0.20 | 0.24 | 0.88 | 2.40 | Download | ---------------------IM-GSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACL-----------FEDVVPMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSRQ- | |||||||||||||||||||
| 2 | 5tgzA | 0.19 | 0.23 | 0.86 | 2.06 | Download | -----------------------SQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDF-HVFHRKDSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLL-------GWNCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKAHSHAPDQARIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKLIKTVFFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 3 | 4iaqA | 0.16 | 0.21 | 0.87 | 1.91 | Download | --------------YIYQDSISLPWKVLLVMLLALITLATTLSNAFVIATVYRTRKLHTPANYLIASLAVTDLLVSILVMPISTMYTVTGRWTLGQVVCDFWLSSDITCCTASIWHLCVIALDRYWAITDAVEYSAKRTPKRAAVMIALVWVFSISISLPPF-----FWRQASE----------------CVVNTDHILYTVYSTVGAFYFPTLLLIALYGRIYVEARSRIMAARERKATKTLGIILGAFIVCWLPFFIISLVMPIHL-----AIFDFFTWLGYLNSLINPIIYTMSNEDFKQAFHKLIRFK-- | |||||||||||||||||||
| 4 | 4ib4 | 0.18 | 0.23 | 0.89 | 1.52 | Download | -------------------EEQGNKLHWAALLILMVIIPTIGGNTLVILAVSLEKKLQYATNYFLMSLAVADLLVGLFVMPIALLTIMFAMWPLPLVLCPAWLFLDVLFSTASIWHLCAISVDRYIAIKKPIQANQYNSRATAFIKITVVWLISIGIAIPVPIKGIE--T----NPNNITCVLTK---------ERFGDFMLFGSLAAFFTPLAIMIVTYFLTIHALQKKAADSNEQRASKVLGIVFFLFLLMWCPFFITNITLVLCDSCQTQMLLEIFVWIGYVSSGVNPLVYTLFNKTFRDAFGRYITCNYR | |||||||||||||||||||
| 5 | 3uonA | 0.17 | 0.18 | 0.87 | 1.13 | Download | ----------------------TFEVVFIVLVAGSLSLVTIIGNILVMVSIKVNRHLQTVNNYFLFSLACADLIIGVFSMNLYTLYTVIGYWPLGPVVCDLWLALDYVVSNASVMNLLIISFDRYFCVTKPLTYPVKRTTKMAGMMIAAAWVLSFILWAPAILFWQFI--VGVRTVEDGECYIQF---------FSNAAVTFGTAIAAFYLPVIIMTVLYWHISRASKSR----REKKVTRTILAILLAFIITWAPYNVMVLINTFCAPCIPNTVWTIGYWLCYINSTINPACYALCNATFKKTFKHLLM---- | |||||||||||||||||||
| 6 | 3emlA | 0.19 | 0.24 | 0.88 | 2.58 | Download | -----------------------MGSSVYITVELAIAVLAILGNVLVCWAVWLNSNLQNVTNYFVVSLAAADIAVGVLAIPFAITIST--GFCAACHGCLFIACFVLVLTQSSIFSLLAIAIDRYIAIRIPLRYNGLVTGTRAKGIIAICWVLSFAIGLTPMLGWNNCGGCGEGQVACLFEDVV-----------PMNYMVYFNFFACVLVPLLLMLGVYLRIFLAARRQLVHAAKSLAIIVGLFALCWLPLHIINCFTFFC---PDCSHAPLWLMYLAIVLSHTNSVVNPFIYAYRIREFRQTFRKIIRSHVL | |||||||||||||||||||
| 7 | 2rh1 | 0.17 | 0.21 | 0.62 | 1.70 | Download | -------------------------VVGMGIVMSLIVLAIVFGNVLVITAIAKFERLQTVTNYFITSLACADLVMGLAVVPFGAAHILMKMWTFGNFWCEFWTSIDVLCVTASIETLCVIAVDRYFAITSPFKYQSLLTKNKARVIILMVWIVSGLTSFLPIQMWYRAT---HQ--EAINCYAE----ETCCDFFTNQAYAIASSIVSFYVPLVIMVFVYSRVFQEAKR------------------------------------------------------------------------------------- | |||||||||||||||||||
| 8 | 4buoA | 0.18 | 0.20 | 0.90 | 2.92 | Download | ----NSDL---------DVNTDIYSKVLVTAIYLALFVVGTVGNSVTLFTLARKKSLQSTVDYYLGSLALSDLLILLLAMPVELYNFIWVHHPWAFAGCRGYYFLRDACTYATALNVVSLSVELYLAICHPFKAKTLMSRSRTKKFISAIWLASALLAIPMLFTMGLQNLSGDGTHPG------------GVDTATLKVVIQVNTFMSFLFPMLVASILNTVIANKLTVMVHQPGRVQALRRGVLVLRAVVIAFVHVRRLMFCYISDEQWTYHYFYMLTNALVYVSAAINPILYNLVSANFRQVFLSTL----- | |||||||||||||||||||
| 9 | 5tgzA | 0.18 | 0.23 | 0.91 | 3.12 | Download | -GGRGENFMDIECFMVLN----PSQQLAIAVLSLTLGTFTVLENLLVLCVILHSRSLRRPSYHFIGSLAVADLLGSVIFVYSFIDFHVFHRK-DSRNVFLFKLGGVTASFTASVGSLFLAAIDRYISIHRPLAYKRIVTRPKAVVAFCLMWTIAIVIAVLPLLG--------NCEKLQSVC---------SDIFPHIDKTYLMFWIGVVSVLLLFIVYAYMYILWKA-APDQARMDIELAKTLVLILVVLIICWGPLLAIMVYDVFGKMNKIKTVFAFCSMLCLLNSTVNPIIYALRSKDLRHAFRSMF----- | |||||||||||||||||||
| 10 | 4gpoA | 0.17 | 0.19 | 0.89 | 4.76 | Download | ---------------------LQQWEAGMSLLMALVVLLIVAGNVLVIAAIGSTQRLQTLTNLFITSLACADLVVGLLVVPFGATLVVRGTWLWGSFLCELWTSLDVLCVTASIETLCVIAIDRYLAITSPFRYQSLMTRARAKVIICTVWAISALVSFLPIMMHWWRDEDPQALKCYQDPG--------CCDFVTNRAYAIASSIISFYIPLLIMIFVALRVYREAKEQVMLMREHKALKTLGIIMGVFTLCWLPFFLVNIVNVFNRDLVPDWLFVAFNWLGYANSAMNPIIYC-RSPDFRKAFKRL------ | |||||||||||||||||||
| ||||||||||||||||||||||||||
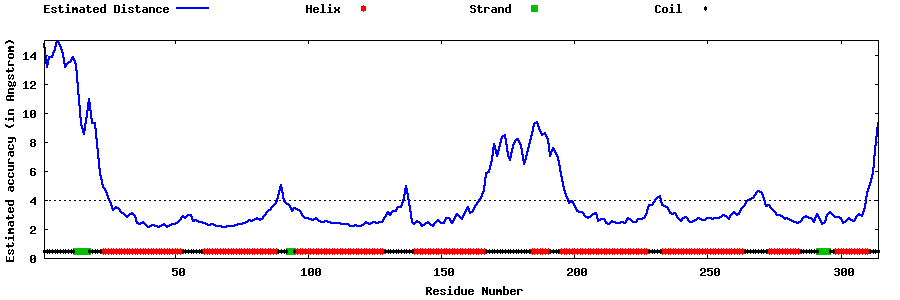
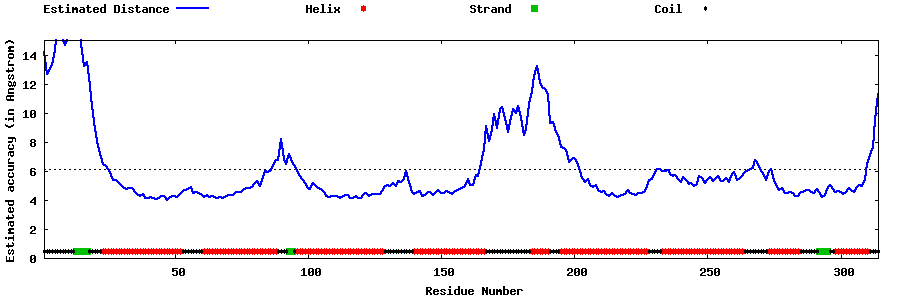
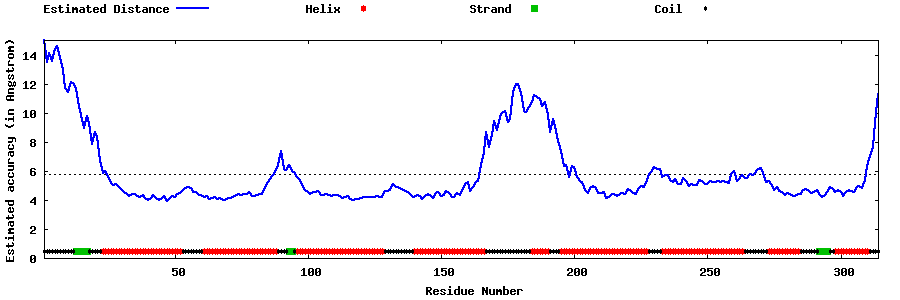
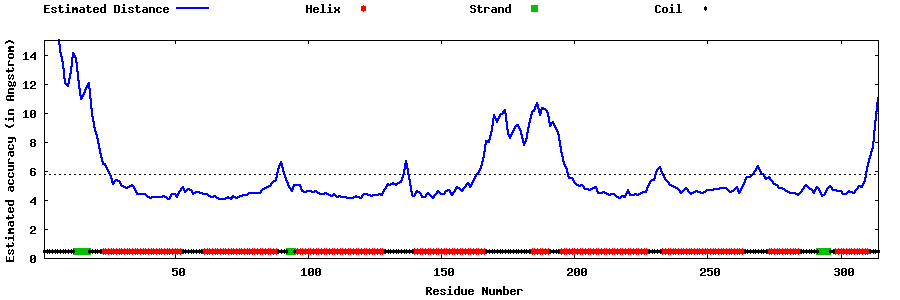
| Generated 3D models | Estimated local accuracy of models | ||
 | |||
|
|
|||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||
 | |||
|
| |||