

You can:
| Name | Proteinase-activated receptor 2 |
|---|---|
| Species | Homo sapiens (Human) |
| Gene | F2RL1 |
| Synonym | Protease-activated receptor-2 PAR2 PAR-2 GPR11 G-protein coupled receptor 11 [ Show all ] |
| Disease | N/A |
| Length | 397 |
| Amino acid sequence | MRSPSAAWLLGAAILLAASLSCSGTIQGTNRSSKGRSLIGKVDGTSHVTGKGVTVETVFSVDEFSASVLTGKLTTVFLPIVYTIVFVVGLPSNGMALWVFLFRTKKKHPAVIYMANLALADLLSVIWFPLKIAYHIHGNNWIYGEALCNVLIGFFYGNMYCSILFMTCLSVQRYWVIVNPMGHSRKKANIAIGISLAIWLLILLVTIPLYVVKQTIFIPALNITTCHDVLPEQLLVGDMFNYFLSLAIGVFLFPAFLTASAYVLMIRMLRSSAMDENSEKKRKRAIKLIVTVLAMYLICFTPSNLLLVVHYFLIKSQGQSHVYALYIVALCLSTLNSCIDPFVYYFVSHDFRDHAKNALLCRSVRTVKQMQVSLTSKKHSRKSSSYSSSSTTVKTSY |
| UniProt | P55085 |
| Protein Data Bank | 5ndz, 5ndd |
| GPCR-HGmod model | P55085 |
| 3D structure model | This structure is from PDB ID 5ndz. |
| BioLiP | BL0377325, BL0377326 |
| Therapeutic Target Database | N/A |
| ChEMBL | CHEMBL5963 |
| IUPHAR | 348 |
| DrugBank | N/A |
| Name | CHEMBL3735017 |
|---|---|
| Molecular formula | C19H12Cl3N3O2 |
| IUPAC name | 6-chloro-2-(2,4-dichlorophenoxy)-N-(furan-2-ylmethyl)quinazolin-4-amine |
| Molecular weight | 420.674 |
| Hydrogen bond acceptor | 5 |
| Hydrogen bond donor | 1 |
| XlogP | 6.2 |
| Synonyms | N/A |
| Inchi Key | GUDINIUSGMZTHX-UHFFFAOYSA-N |
| Inchi ID | InChI=1S/C19H12Cl3N3O2/c20-11-3-5-16-14(8-11)18(23-10-13-2-1-7-26-13)25-19(24-16)27-17-6-4-12(21)9-15(17)22/h1-9H,10H2,(H,23,24,25) |
| PubChem CID | 127036507 |
| ChEMBL | CHEMBL3735017 |
| IUPHAR | N/A |
| BindingDB | N/A |
| DrugBank | N/A |
Structure | 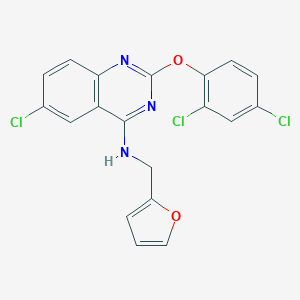 |
| Lipinski's druglikeness | This ligand has a partition coefficient log P greater than 5. |
| Parameter | Value | Reference | Database source |
|---|---|---|---|
| Inhibition | >10.0 % | PMID26631441 | ChEMBL |
zhanglab![]() zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417
zhanggroup.org | +65-6601-1241 | Computing 1, 13 Computing Drive, Singapore 117417